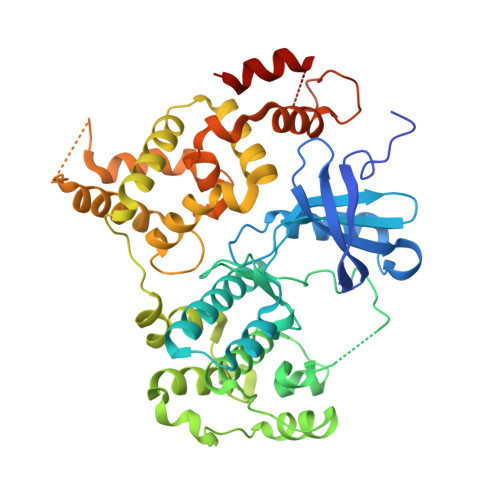
Discovery of Potent and Selective Inhibitors of Calcium-Dependent Protein Kinase 1 (CDPK1) from C. parvum and T. gondii.
Murphy, R.C., Ojo, K.K., Larson, E.T., Castellanos-Gonzalez, A., Perera, B.G., Keyloun, K.R., Kim, J.E., Bhandari, J.G., Muller, N.R., Verlinde, C.L., White, A.C., Merritt, E.A., Van Voorhis, W.C., Maly, D.J.(2010) ACS Med Chem Lett 1: 331-335
- PubMed: 21116453 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml100096t
- Primary Citation Related Structures:
3MWU, 3N51, 3NCG - PubMed Abstract:
The protozoans Cryptosporidium parvum and Toxoplasma gondii are parasites of major health concern to humans. Both parasites contain a group of calcium-dependent protein kinases (CDPKs), which are found in plants and ciliates but not in humans or fungi. Here we describe a series of potent inhibitors that target CDPK1 in C. parvum (CpCDPK1) and T. gondii (TgCDPK1). These inhibitors are highly selective for CpCDPK1 and TgCDPK1 over the mammalian kinases SRC and ABL. Furthermore, they are able to block an early stage of C. parvum invasion of HCT-8 host cells, which is similar to their effects on T. gondii invasion of human fibroblasts.
- Department of Chemistry, University of Washington.
Organizational Affiliation: