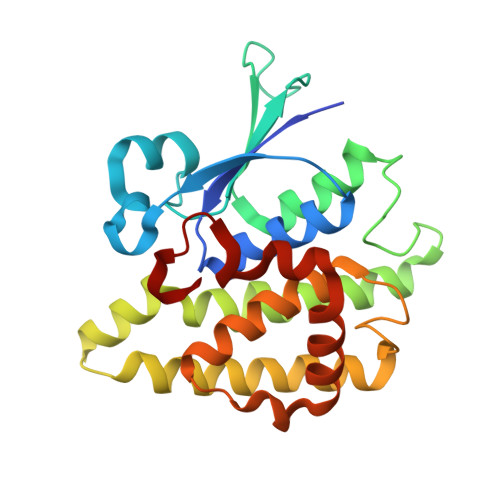
Structures of a putative zeta-class glutathione S-transferase from the pathogenic fungus Coccidioides immitis.
Edwards, T.E., Bryan, C.M., Leibly, D.J., Dieterich, S.H., Abendroth, J., Sankaran, B., Sivam, D., Staker, B.L., Van Voorhis, W.C., Myler, P.J., Stewart, L.J.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1038-1043
- PubMed: 21904047 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111009493
- Primary Citation Related Structures:
3LG6, 3N5O - PubMed Abstract:
Coccidioides immitis is a pathogenic fungus populating the southwestern United States and is a causative agent of coccidioidomycosis, sometimes referred to as Valley Fever. Although the genome of this fungus has been sequenced, many operons are not properly annotated. Crystal structures are presented for a putative uncharacterized protein that shares sequence similarity with ζ-class glutathione S-transferases (GSTs) in both apo and glutathione-bound forms. The apo structure reveals a nonsymmetric homodimer with each protomer comprising two subdomains: a C-terminal helical domain and an N-terminal thioredoxin-like domain that is common to all GSTs. Half-site binding is observed in the glutathione-bound form. Considerable movement of some components of the active site relative to the glutathione-free form was observed, indicating an induced-fit mechanism for cofactor binding. The sequence homology, structure and half-site occupancy imply that the protein is a ζ-class glutathione S-transferase, a maleylacetoacetate isomerase (MAAI).
- Seattle Structural Genomics Center for Infectious Disease, USA. tedwards@embios.com
Organizational Affiliation: