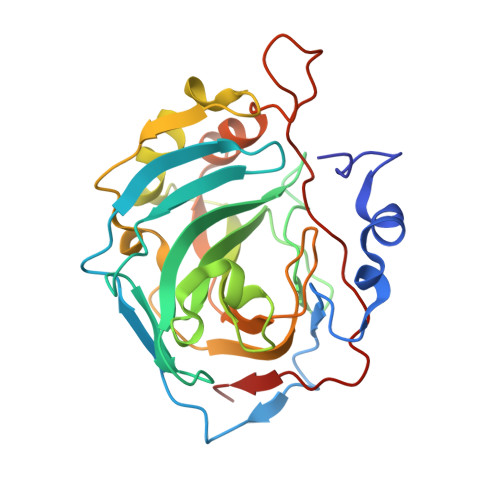
Structural and kinetic study of the extended active site for proton transfer in human carbonic anhydrase II.
Domsic, J.F., Williams, W., Fisher, S.Z., Tu, C., Agbandje-McKenna, M., Silverman, D.N., McKenna, R.(2010) Biochemistry 49: 6394-6399
- PubMed: 20578724 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi1007645
- Primary Citation Related Structures:
3MNH, 3MNI, 3MNJ, 3MNK - PubMed Abstract:
The catalysis of CO(2) hydration by human carbonic anhydrase II (HCA II) is limited in maximal velocity by proton transfer from a zinc-bound water molecule to the proton shuttle His64. This proton transfer occurs along a hydrogen-bonded water network, leading to the proton shuttle residue His64, which in turn transfers the proton to bulk solvent. The side chain of His64 occupies two conformations in wild-type HCA II, pointing inward toward the zinc or outward toward bulk solvent. Previously, several studies have examined the roles of residues of the active site cavity that interact with the solvent-mediated hydrogen-bonded network between His64 and the zinc-bound water. Here these studies are extended to examine the effects on proton transfer by mutation at Lys170 (to Ala, Asp, Glu, and His), a residue located near the side chain of His64 but over 15 A away from the active site zinc. In all four variants, His64 is observed in the inward conformation associated with a decrease in the pK(a) of His64 by as much as 1.0 unit and an increase in the rate constant for proton transfer to as much as 4 micros(-1), approximately 5-fold larger than wild-type HCA II. The results show a significant extension of the effective active site of HCA II from the zinc-bound water at the base of the conical cavity in the enzyme to Lys170 near the rim of the cavity. These data emphasize that the active site of HCA II is extended to include residues that, at first glance, appear to be too far from the zinc to exert any catalytic effects.
- Department of Biochemistry and Molecular Biology, University of Florida, Gainesville, FL 32610, USA.
Organizational Affiliation: