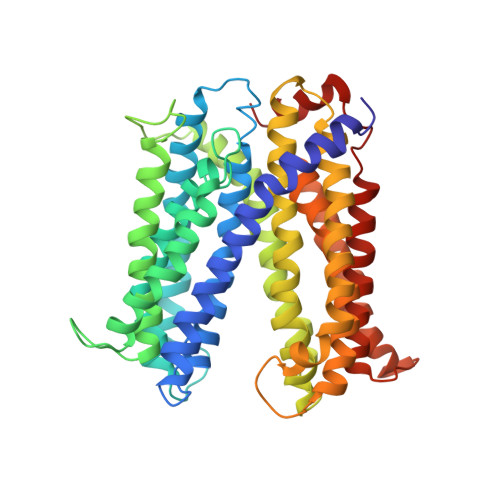
Structure of a cation-bound multidrug and toxic compound extrusion transporter.
He, X., Szewczyk, P., Karyakin, A., Evin, M., Hong, W.X., Zhang, Q., Chang, G.(2010) Nature 467: 991-994
- PubMed: 20861838 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09408
- Primary Citation Related Structures:
3MKT, 3MKU - PubMed Abstract:
Transporter proteins from the MATE (multidrug and toxic compound extrusion) family are vital in metabolite transport in plants, directly affecting crop yields worldwide. MATE transporters also mediate multiple-drug resistance (MDR) in bacteria and mammals, modulating the efficacy of many pharmaceutical drugs used in the treatment of a variety of diseases. MATE transporters couple substrate transport to electrochemical gradients and are the only remaining class of MDR transporters whose structure has not been determined. Here we report the X-ray structure of the MATE transporter NorM from Vibrio cholerae determined to 3.65 Å, revealing an outward-facing conformation with two portals open to the outer leaflet of the membrane and a unique topology of the predicted 12 transmembrane helices distinct from any other known MDR transporter. We also report a cation-binding site in close proximity to residues previously deemed critical for transport. This conformation probably represents a stage of the transport cycle with high affinity for monovalent cations and low affinity for substrates.
- Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, CB105, La Jolla, California 92037, USA.
Organizational Affiliation: