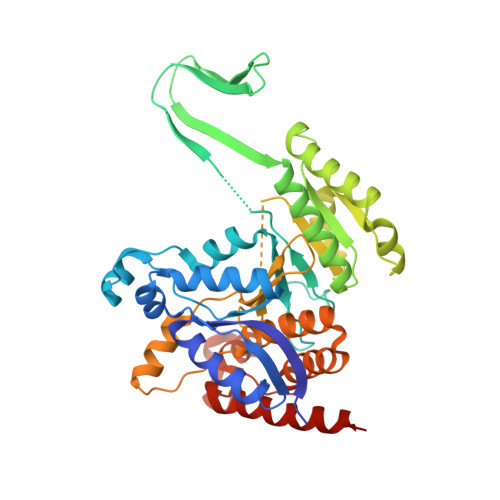
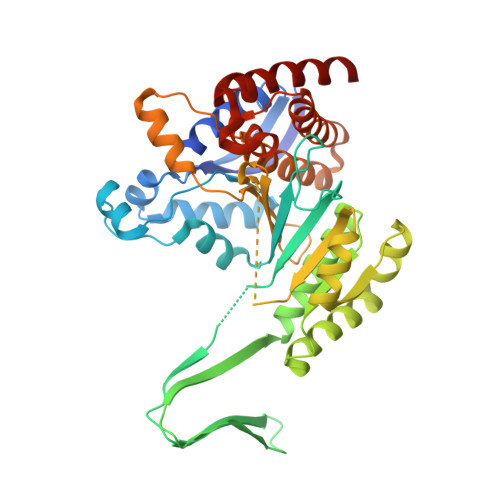
Molecular mechanisms of "off-on switch" of activities of human IDH1 by tumor-associated mutation R132H.
Yang, B., Zhong, C., Peng, Y., Lai, Z., Ding, J.(2010) Cell Res 20: 1188-1200
- PubMed: 20975740 Search on PubMed
- DOI: https://doi.org/10.1038/cr.2010.145
- Primary Citation Related Structures:
3MAP, 3MAR, 3MAS - PubMed Abstract:
Human cytosolic NADP-IDH (IDH1) has recently been found to be involved in tumorigenesis. Notably, the tumor-derived IDH1 mutations identified so far mainly occur at Arg132, and mutation R132H is the most prevalent one. This mutation impairs the oxidative IDH activity of the enzyme, but renders a new reduction function of converting α-ketoglutarate (αKG) to 2-hydroxyglutarate. Here, we report the structures of the R132H mutant IDH1 with and without isocitrate (ICT) bound. The structural data together with mutagenesis and biochemical data reveal a previously undefined initial ICT-binding state and demonstrate that IDH activity requires a conformational change to a closed pre-transition state. Arg132 plays multiple functional roles in the catalytic reaction; in particular, the R132H mutation hinders the conformational changes from the initial ICT-binding state to the pre-transition state, leading to the impairment of the IDH activity. Our results describe for the first time that there is an intermediate conformation that corresponds to an initial ICT-binding state and that the R132H mutation can trap the enzyme in this conformation, therefore shedding light on the molecular mechanism of the "off switch" of the potentially tumor-suppressive IDH activity. Furthermore, we proved the necessity of Tyr139 for the gained αKG reduction activity and propose that Tyr139 may play a vital role by compensating the increased negative charge on the C2 atom of αKG during the transfer of a hydride anion from NADPH to αKG, which provides new insights into the mechanism of the "on switch" of the hypothetically oncogenic reduction activity of IDH1 by this mutation.
- State Key Laboratory of Molecular Biology and Research Center for Structural Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai 200031, China.
Organizational Affiliation: