Quantitative analysis of processive RNA degradation by the archaeal RNA exosome
Hartung, S., Niederberger, T., Hartung, M., Tresch, A., Hopfner, K.-P.(2010) Nucleic Acids Res 38: 5166-5176
- PubMed: 20392821
- DOI: https://doi.org/10.1093/nar/gkq238
- Primary Citation of Related Structures:
3M7N, 3M85 - PubMed Abstract:
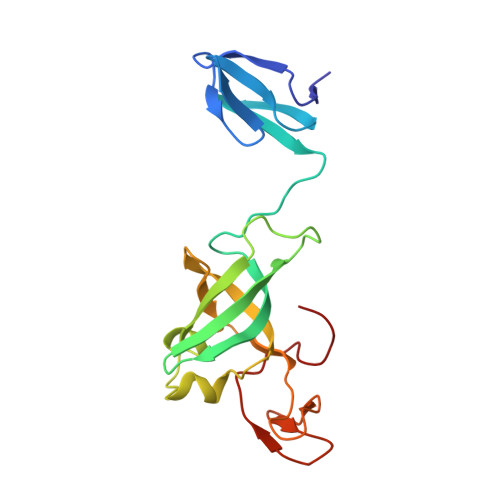
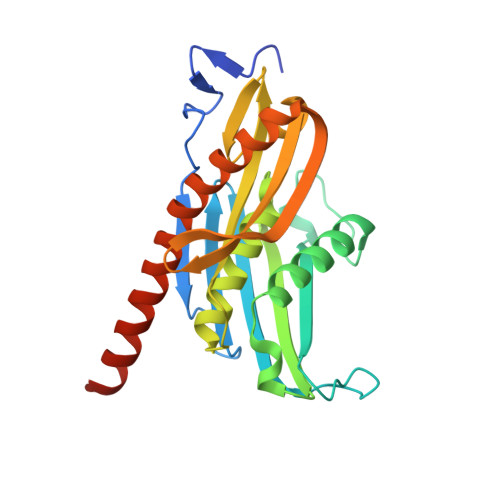
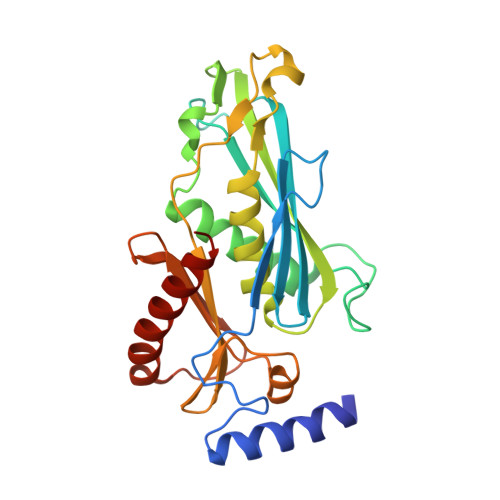
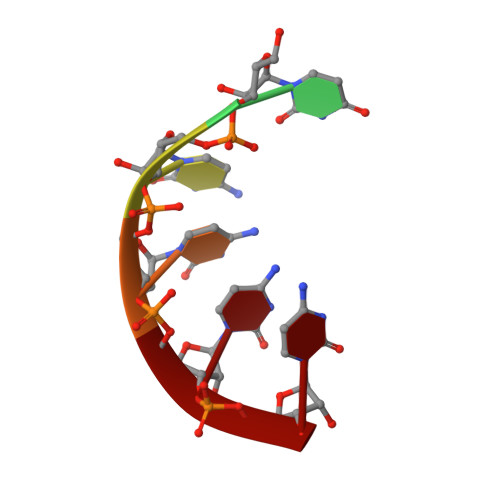
RNA exosomes are large multisubunit assemblies involved in controlled RNA processing. The archaeal exosome possesses a heterohexameric processing chamber with three RNase-PH-like active sites, capped by Rrp4- or Csl4-type subunits containing RNA-binding domains. RNA degradation by RNA exosomes has not been studied in a quantitative manner because of the complex kinetics involved, and exosome features contributing to efficient RNA degradation remain unclear. Here we derive a quantitative kinetic model for degradation of a model substrate by the archaeal exosome. Markov Chain Monte Carlo methods for parameter estimation allow for the comparison of reaction kinetics between different exosome variants and substrates. We show that long substrates are degraded in a processive and short RNA in a more distributive manner and that the cap proteins influence degradation speed. Our results, supported by small angle X-ray scattering, suggest that the Rrp4-type cap efficiently recruits RNA but prevents fast RNA degradation of longer RNAs by molecular friction, likely by RNA contacts to its unique KH-domain. We also show that formation of the RNase-PH like ring with entrapped RNA is not required for high catalytic efficiency, suggesting that the exosome chamber evolved for controlled processivity, rather than for catalytic chemistry in RNA decay.
- Center for Integrated Protein Sciences, Department of Biochemistry, Ludwig-Maximilians-University Munich, Munich, Germany.
Organizational Affiliation: