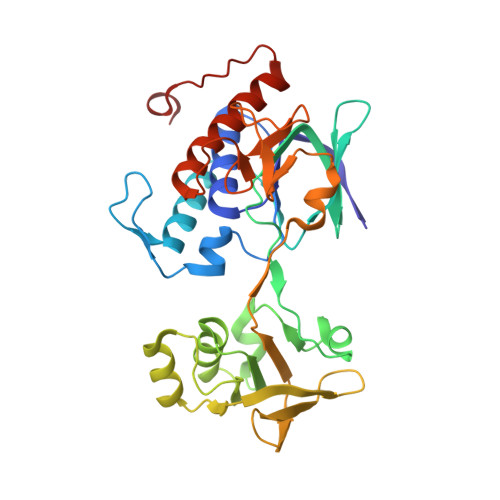
Crystal structure analysis of Bacillus subtilis ferredoxin-NADP(+) oxidoreductase and the structural basis for its substrate selectivity
Komori, H., Seo, D., Sakurai, T., Higuchi, Y.(2010) Protein Sci 19: 2279-2290
- PubMed: 20878669 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.508
- Primary Citation Related Structures:
3LZW, 3LZX - PubMed Abstract:
Bacillus subtilis yumC encodes a novel type of ferredoxin-NADP+ oxidoreductase (FNR) with a primary sequence and oligomeric conformation distinct from those of previously known FNRs. In this study, the crystal structure of B. subtilis FNR (BsFNR) complexed with NADP+ has been determined. BsFNR features two distinct binding domains for FAD and NADPH in accordance with its structural similarity to Escherichia coli NADPH-thioredoxin reductase (TdR) and TdR-like protein from Thermus thermophilus HB8 (PDB code: 2ZBW). The deduced mode of NADP+ binding to the BsFNR molecule is nonproductive in that the nicotinamide and isoalloxazine rings are over 15 Å apart. A unique C-terminal extension, not found in E. coli TdR but in TdR-like protein from T. thermophilus HB8, covers the re-face of the isoalloxazine moiety of FAD. In particular, Tyr50 in the FAD-binding region and His324 in the C-terminal extension stack on the si- and re-faces of the isoalloxazine ring of FAD, respectively. Aromatic residues corresponding to Tyr50 and His324 are also found in the plastid-type FNR superfamily of enzymes, and the residue corresponding to His324 has been reported to be responsible for nucleotide specificity. In contrast to the plastid-type FNRs, replacement of His324 with Phe or Ser had little effect on the specificity or reactivity of BsFNR with NAD(P)H, whereas replacement of Arg190, which interacts with the 2'-phosphate of NADP+, drastically decreased its affinity toward NADPH. This implies that BsFNR adopts the same nucleotide binding mode as the TdR enzyme family and that aromatic residue on the re-face of FAD is hardly relevant to the nucleotide selectivity.
- Department of Life Science, Graduate School of Life Science, University of Hyogo, Ako-gun, Hyogo 678-1297, Japan.
Organizational Affiliation: