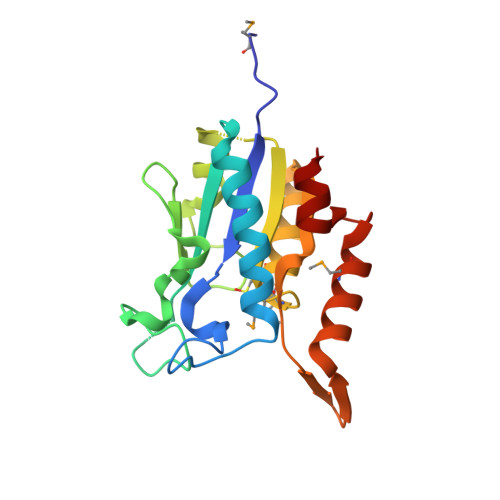
Crystal structure of a putative isochorismatase hydrolase from Oleispira antarctica.
Goral, A.M., Tkaczuk, K.L., Chruszcz, M., Kagan, O., Savchenko, A., Minor, W.(2012) J Struct Funct Genomics 13: 27-36
- PubMed: 22350524
- DOI: https://doi.org/10.1007/s10969-012-9127-5
- Primary Citation of Related Structures:
3LQY - PubMed Abstract:
Isochorismatase-like hydrolases (IHL) constitute a large family of enzymes divided into five structural families (by SCOP). IHLs are crucial for siderophore-mediated ferric iron acquisition by cells. Knowledge of the structural characteristics of these molecules will enhance the understanding of the molecular basis of iron transport, and perhaps resolve which of the mechanisms previously proposed in the literature is the correct one. We determined the crystal structure of the apo-form of a putative isochorismatase hydrolase OaIHL (PDB code: 3LQY) from the antarctic γ-proteobacterium Oleispira antarctica, and did comparative sequential and structural analysis of its closest homologs. The characteristic features of all analyzed structures were identified and discussed. We also docked isochorismate to the determined crystal structure by in silico methods, to highlight the interactions of the active center with the substrate. The putative isochorismate hydrolase OaIHL from O. antarctica possesses the typical catalytic triad for IHL proteins. Its active center resembles those IHLs with a D-K-C catalytic triad, rather than those variants with a D-K-X triad. OaIHL shares some structural and sequential features with other members of the IHL superfamily. In silico docking results showed that despite small differences in active site composition, isochorismate binds to in the structure of OaIHL in a similar mode to its binding in phenazine biosynthesis protein PhzD (PDB code 1NF8).
- Department of Molecular Physiology and Biological Physics, University of Virginia, 1340 Jefferson Park Avenue, Charlottesville, VA 22908, USA.
Organizational Affiliation: