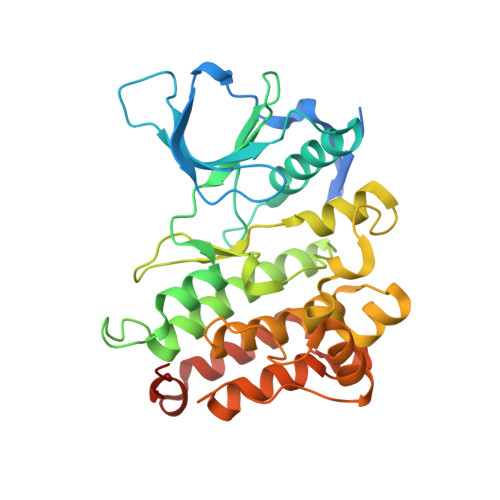
Crystal structure of the ALK (anaplastic lymphoma kinase) catalytic domain.
Lee, C.C., Jia, Y., Li, N., Sun, X., Ng, K., Ambing, E., Gao, M.Y., Hua, S., Chen, C., Kim, S., Michellys, P.Y., Lesley, S.A., Harris, J.L., Spraggon, G.(2010) Biochem J 430: 425-437
- PubMed: 20632993 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20100609
- Primary Citation Related Structures:
3L9P, 3LCS, 3LCT - PubMed Abstract:
ALK (anaplastic lymphoma kinase) is an RTK (receptor tyrosine kinase) of the IRK (insulin receptor kinase) superfamily, which share an YXXXYY autophosphorylation motif within their A-loops (activation loops). A common activation and regulatory mechanism is believed to exist for members of this superfamily typified by IRK and IGF1RK (insulin-like growth factor receptor kinase-1). Chromosomal translocations involving ALK were first identified in anaplastic large-cell lymphoma, a subtype of non-Hodgkin's lymphoma, where aberrant fusion of the ALK kinase domain with the NPM (nucleophosmin) dimerization domain results in autophosphosphorylation and ligand-independent activation. Activating mutations within the full-length ALK kinase domain, most commonly R1275Q and F1174L, which play a major role in neuroblastoma, were recently identified. To provide a structural framework for understanding these mutations and to guide structure-assisted drug discovery efforts, the X-ray crystal structure of the unphosphorylated ALK catalytic domain was determined in the apo, ADP- and staurosporine-bound forms. The structures reveal a partially inactive protein kinase conformation distinct from, and lacking, many of the negative regulatory features observed in inactive IGF1RK/IRK structures in their unphosphorylated forms. The A-loop adopts an inhibitory pose where a short proximal A-loop helix (alphaAL) packs against the alphaC helix and a novel N-terminal beta-turn motif, whereas the distal portion obstructs part of the predicted peptide-binding region. The structure helps explain the reported unique peptide substrate specificity and the importance of phosphorylation of the first A-loop Tyr1278 for kinase activity and NPM-ALK transforming potential. A single amino acid difference in the ALK substrate peptide binding P-1 site (where the P-site is the phosphoacceptor site) was identified that, in conjunction with A-loop sequence variation including the RAS (Arg-Ala-Ser)-motif, rationalizes the difference in the A-loop tyrosine autophosphorylation preference between ALK and IGF1RK/IRK. Enzymatic analysis of recombinant R1275Q and F1174L ALK mutant catalytic domains confirms the enhanced activity and transforming potential of these mutants. The transforming ability of the full-length ALK mutants in soft agar colony growth assays corroborates these findings. The availability of a three-dimensional structure for ALK will facilitate future structure-function and rational drug design efforts targeting this receptor tyrosine kinase.
- The Genomics Institute of the Novartis Research Foundation, San Diego, CA 92121, USA. clee@gnf.org
Organizational Affiliation: