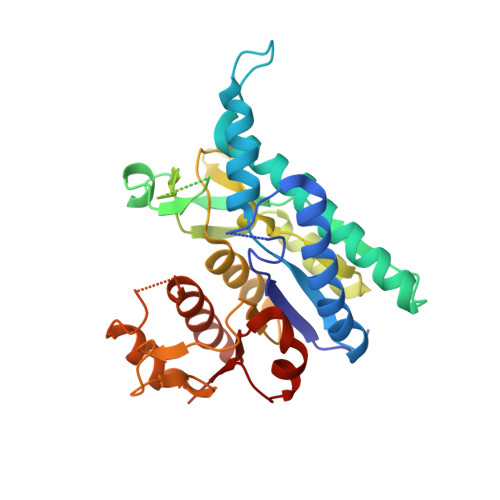
X-ray structure and characterization of carbamate kinase from the human parasite Giardia lamblia.
Galkin, A., Kulakova, L., Wu, R., Nash, T.E., Dunaway-Mariano, D., Herzberg, O.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 386-390
- PubMed: 20383005
- DOI: https://doi.org/10.1107/S1744309110004665
- Primary Citation Related Structures:
3KZF - PubMed Abstract:
Carbamate kinase catalyzes the reversible conversion of carbamoyl phosphate and ADP to ATP and ammonium carbamate, which is hydrolyzed to ammonia and carbonate. The three-dimensional structure of carbamate kinase from the human parasite Giardia lamblia (glCK) has been determined at 3 A resolution. The crystals belonged to the monoclinic space group P2(1), with unit-cell parameters a = 69.77, b = 85.41, c = 102.1 A, beta = 106.8 degrees . The structure was refined to a final R factor of 0.227. The essentiality of glCK together with its absence in humans makes the enzyme an attractive candidate for anti-Giardia drug development. Steady-state kinetic rate constants have been determined. The k(cat) for ATP formation is 319 +/- 9 s(-1). The K(m) values for carbamoyl phosphate and ADP are 85 +/- 6 and 70 +/- 5 microM, respectively. The structure suggests that three invariant lysine residues (Lys131, Lys216 and Lys278) may be involved in the binding of substrates and phosphoryl transfer. The structure of glCK reveals that a glycerol molecule binds in the likely carbamoyl phosphate-binding site.
- W. M. Keck Laboratory for Structural Biology, Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, Rockville, Maryland, USA.
Organizational Affiliation: