Novel synthesis and structural characterization of a high-affinity paramagnetic kinase probe for the identification of non-ATP site binders by nuclear magnetic resonance.
Moy, F.J., Lee, A., Gavrin, L.K., Xu, Z.B., Sievers, A., Kieras, E., Stochaj, W., Mosyak, L., McKew, J., Tsao, D.H.(2010) J Med Chem 53: 1238-1249
- PubMed: 20038108 Search on PubMed
- DOI: https://doi.org/10.1021/jm901525b
- Primary Citation Related Structures:
3KXZ - PubMed Abstract:
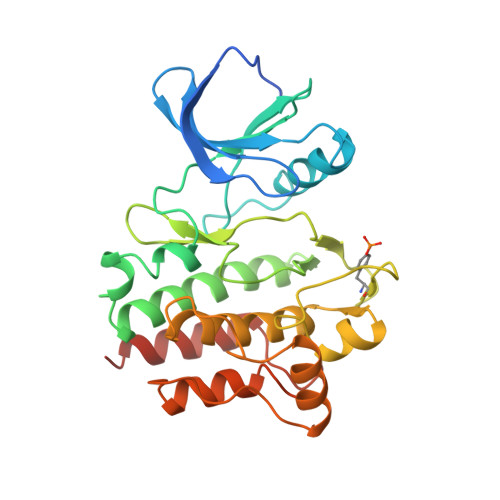
To aid in the pursuit of selective kinase inhibitors, we have developed a unique ATP site binder tool for the detection of binders outside the ATP site by nuclear magnetic resonance (NMR). We report here the novel synthesis that led to this paramagnetic spin-labeled pyrazolopyrimidine probe (1), which exhibits nanomolar inhibitory activity against multiple kinases. We demonstrate the application of this probe by performing NMR binding experiments with Lck and Src kinases and utilize it to detect the binding of two compounds proximal to the ATP site. The complex structure of the probe with Lck is also presented, revealing how the probe fits in the ATP site and the specific interactions it has with the protein. We believe that this spin-labeled probe is a valuable tool that holds broad applicability in a screen for non-ATP site binders.
- Structural Biology and Computational Chemistry, Wyeth Research, 200 CambridgePark Drive, Cambridge, Massachusetts 02140, USA.
Organizational Affiliation: