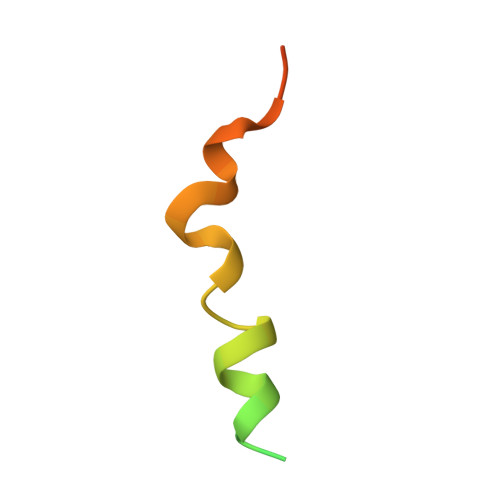
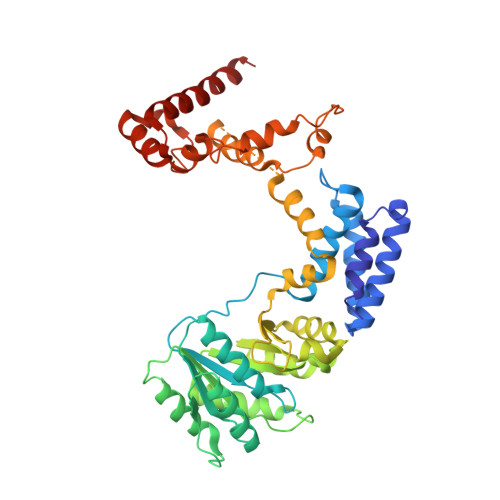
Recognition of a signal peptide by the signal recognition particle.
Janda, C.Y., Li, J., Oubridge, C., Hernandez, H., Robinson, C.V., Nagai, K.(2010) Nature 465: 507-510
- PubMed: 20364120 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature08870
- Primary Citation Related Structures:
3KL4 - PubMed Abstract:
Targeting of proteins to appropriate subcellular compartments is a crucial process in all living cells. Secretory and membrane proteins usually contain an amino-terminal signal peptide, which is recognized by the signal recognition particle (SRP) when nascent polypeptide chains emerge from the ribosome. The SRP-ribosome nascent chain complex is then targeted through its GTP-dependent interaction with SRP receptor to the protein-conducting channel on endoplasmic reticulum membrane in eukaryotes or plasma membrane in bacteria. A universally conserved component of SRP (refs 1, 2), SRP54 or its bacterial homologue, fifty-four homologue (Ffh), binds the signal peptides, which have a highly divergent sequence divisible into a positively charged n-region, an h-region commonly containing 8-20 hydrophobic residues and a polar c-region. No structure has been reported that exemplifies SRP54 binding of any signal sequence. Here we have produced a fusion protein between Sulfolobus solfataricus SRP54 (Ffh) and a signal peptide connected via a flexible linker. This fusion protein oligomerizes in solution through interaction between the SRP54 and signal peptide moieties belonging to different chains, and it is functional, as demonstrated by its ability to bind SRP RNA and SRP receptor FtsY. We present the crystal structure at 3.5 A resolution of an SRP54-signal peptide complex in the dimer, which reveals how a signal sequence is recognized by SRP54.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 0QH, UK.
Organizational Affiliation: