Double trouble-Buffer selection and His-tag presence may be responsible for nonreproducibility of biomedical experiments.
Majorek, K.A., Kuhn, M.L., Chruszcz, M., Anderson, W.F., Minor, W.(2014) Protein Sci 23: 1359-1368
- PubMed: 25044180
- DOI: https://doi.org/10.1002/pro.2520
- Primary Citation Related Structures:
3KKW, 4M3S - PubMed Abstract:
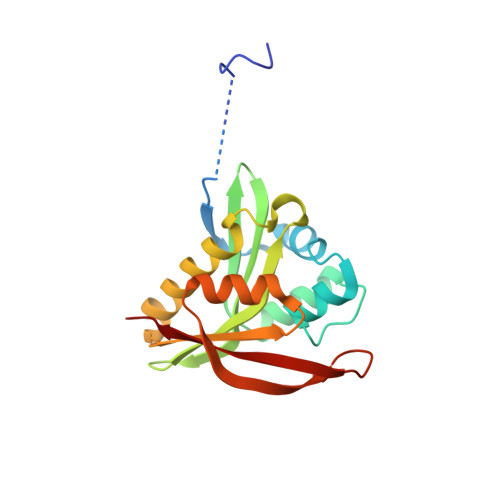
The availability of purified and active protein is the starting point for the majority of in vitro biomedical, biochemical, and drug discovery experiments. The use of polyhistidine affinity tags has resulted in great increases of the efficiency of the protein purification process, but can negatively affect structure and/or activity measurements. Similarly, buffer molecules may perturb the conformational stability of a protein or its activity. During the determination of the structure of a Gcn5-related N-acetyltransferase (GNAT) from Pseudomonas aeruginosa (PA4794), we found that both HEPES and the polyhistidine affinity tag bind (separately) in the substrate-binding site. In the case of HEPES, the molecule induces conformational changes in the active site, but does not significantly affect enzyme activity. In contrast, the uncleaved His-tag does not induce major conformational changes but acts as a weak competitive inhibitor of peptide substrate. In two other GNAT enzymes, we observed that the presence of the His-tag had a strong influence on the activity of these proteins. The influence of protein preparation on functional studies may affect the reproducibility of experiments in other laboratories, even when changes between protocols seem at first glance to be insignificant. Moreover, the results presented here show how critical it is to adjust the experimental conditions for each protein or family of proteins, and investigate the influence of these factors on protein activity and structure, as they may significantly alter the effectiveness of functional characterization and screening methods. Thus, we show that a polyhistidine tag and the buffer molecule HEPES bind in the substrate-binding site and influence the conformation of the active site and the activity of GNAT acetyltransferases. We believe that such discrepancies can influence the reproducibility of some experiments and therefore could have a significant "ripple effect" on subsequent studies.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, Virginia, 22908; Bioinformatics Laboratory, Institute of Molecular Biology and Biotechnology, Faculty of Biology, Adam Mickiewicz University, 61-614, Poznan, Poland; Midwest Center for Structural Genomics, USA; Center for Structural Genomics of Infectious Diseases (CSGID), USA.
Organizational Affiliation: