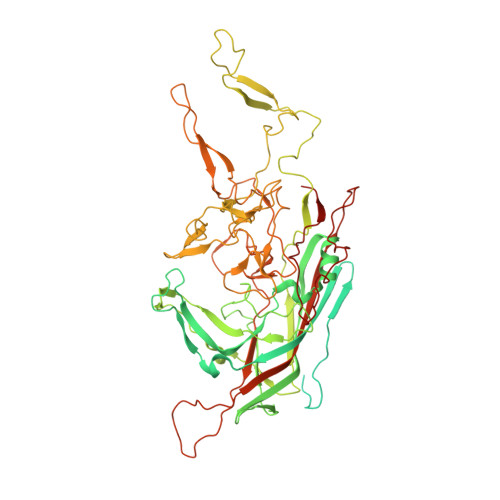
The structure of adeno-associated virus serotype 3B (AAV-3B): insights into receptor binding and immune evasion.
Lerch, T.F., Xie, Q., Chapman, M.S.(2010) Virology 403: 26-36
- PubMed: 20444480 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.virol.2010.03.027
- Primary Citation Related Structures:
3KIC, 3KIE - PubMed Abstract:
Adeno-associated viruses (AAVs) are leading candidate vectors for human gene therapy. AAV serotypes have broad cellular tropism and use a variety of cellular receptors. AAV serotype 3 binds to heparan sulfate proteoglycan prior to cell entry and is serologically distinct from other serotypes. The capsid features that distinguish AAV-3B from other serotypes are poorly understood. The structure of AAV-3B has been determined to 2.6A resolution from twinned crystals of an infectious virus. The most distinctive structural features are located in regions implicated in receptor and antibody binding, providing insights into the cell entry mechanisms and antigenic nature of AAVs. We show that AAV-3B has a lower affinity for heparin than AAV-2, which can be rationalized by the distinct features of the AAV-3B capsid. The structure of AAV-3B provides an additional foundation for the future engineering of improved gene therapy vectors with modified receptor binding or antigenic characteristics.
- Department of Biochemistry and Molecular Biology, School of Medicine, Oregon Health & Science University, Portland, OR 97239-3098, USA.
Organizational Affiliation: