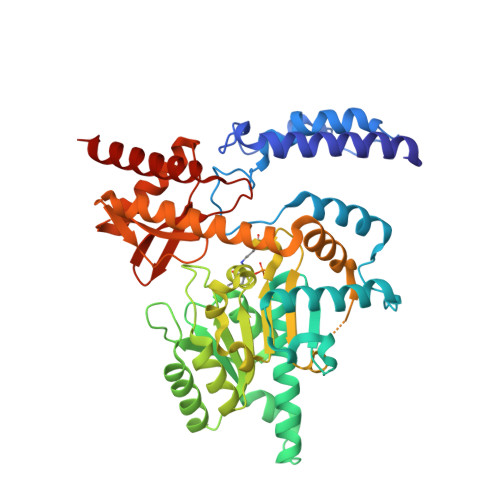
Crystal structure and substrate specificity of Drosophila 3,4-dihydroxyphenylalanine decarboxylase
Han, Q., Ding, H., Robinson, H., Christensen, B.M., Li, J.(2010) PLoS One 5: e8826-e8826
- PubMed: 20098687 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0008826
- Primary Citation Related Structures:
3K40 - PubMed Abstract:
3,4-Dihydroxyphenylalanine decarboxylase (DDC), also known as aromatic L-amino acid decarboxylase, catalyzes the decarboxylation of a number of aromatic L-amino acids. Physiologically, DDC is responsible for the production of dopamine and serotonin through the decarboxylation of 3,4-dihydroxyphenylalanine and 5-hydroxytryptophan, respectively. In insects, both dopamine and serotonin serve as classical neurotransmitters, neuromodulators, or neurohormones, and dopamine is also involved in insect cuticle formation, eggshell hardening, and immune responses. In this study, we expressed a typical DDC enzyme from Drosophila melanogaster, critically analyzed its substrate specificity and biochemical properties, determined its crystal structure at 1.75 Angstrom resolution, and evaluated the roles residues T82 and H192 play in substrate binding and enzyme catalysis through site-directed mutagenesis of the enzyme. Our results establish that this DDC functions exclusively on the production of dopamine and serotonin, with no activity to tyrosine or tryptophan and catalyzes the formation of serotonin more efficiently than dopamine. The crystal structure of Drosophila DDC and the site-directed mutagenesis study of the enzyme demonstrate that T82 is involved in substrate binding and that H192 is used not only for substrate interaction, but for cofactor binding of drDDC as well. Through comparative analysis, the results also provide insight into the structure-function relationship of other insect DDC-like proteins.
- Department of Biochemistry, Virginia Tech, Blacksburg, Virginia, United States of America.
Organizational Affiliation: