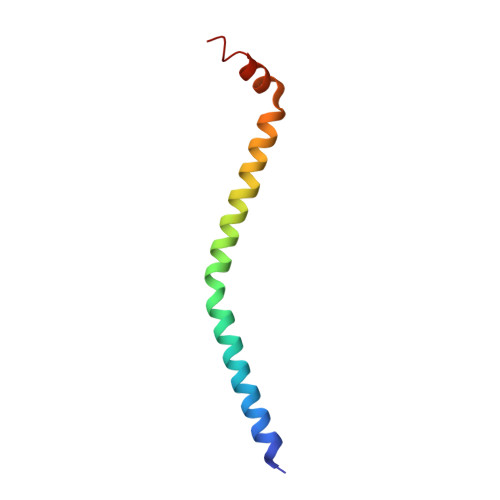
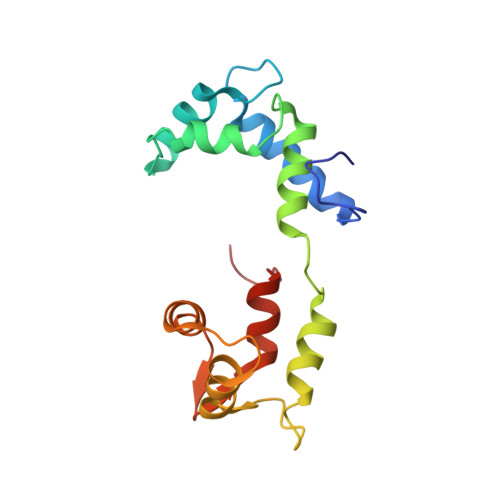
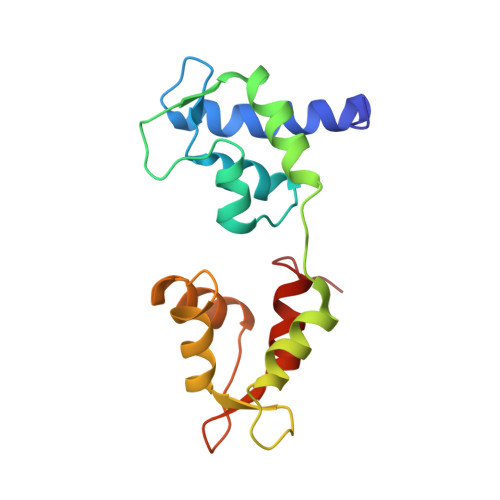
The on-off switch in regulated myosins: different triggers but related mechanisms.
Himmel, D.M., Mui, S., O'Neall-Hennessey, E., Szent-Gyorgyi, A.G., Cohen, C.(2009) J Mol Biology 394: 496-505
- PubMed: 19769984
- DOI: https://doi.org/10.1016/j.jmb.2009.09.035
- Primary Citation Related Structures:
3JTD, 3JVT - PubMed Abstract:
In regulated myosin, motor and enzymatic activities are toggled between the on-state and off-state by a switch located on its lever arm domain, here called the regulatory domain (RD). This region consists of a long alpha-helical "heavy chain" stabilized by a "regulatory" light chain (RLC) and an "essential" light chain (ELC). The on-state is activated by phosphorylation of the RLC of vertebrate smooth muscle RD or by direct binding of Ca(2+) to the ELC of molluscan RD. Crystal structures are available only for the molluscan RD. To understand in more detail the pathway between the on-state and the off-state, we have now also determined the crystal structure of a molluscan (scallop) RD in the absence of Ca(2+). Our results indicate that loss of Ca(2+) abolishes most of the interactions between the light chains and may increase the flexibility of the RD heavy chain. We propose that disruption of critical links with the C-lobe of the RLC is the key event initiating the off-state in both smooth muscle myosins and molluscan myosins.
- Rosenstiel Basic Medical Sciences Research Center, Biology Department, Brandeis University, Waltham, MA 02453-2728, USA. himmel@cabm.rutgers.edu
Organizational Affiliation: