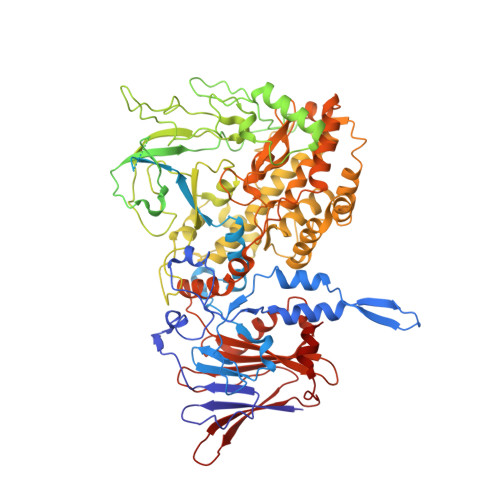
Atomic model of a nonenveloped virus reveals pH sensors for a coordinated process of cell entry.
Zhang, X., Patel, A., Celma, C.C., Yu, X., Roy, P., Zhou, Z.H.(2016) Nat Struct Mol Biol 23: 74-80
- PubMed: 26641711
- DOI: https://doi.org/10.1038/nsmb.3134
- Primary Citation of Related Structures:
3J9D, 3J9E - PubMed Abstract:
Viruses sense environmental cues such as pH to engage in membrane interactions for cell entry during infection, but how nonenveloped viruses sense pH is largely undefined. Here, we report both high- and low-pH structures of bluetongue virus (BTV), which enters cells via a two-stage endosomal process. The receptor-binding protein VP2 possesses a zinc finger that may function to maintain VP2 in a metastable state and a conserved His866, which senses early-endosomal pH. The membrane-penetration protein VP5 has three domains: dagger, unfurling and anchoring. Notably, the β-meander motif of the anchoring domain contains a histidine cluster that can sense late-endosomal pH and also possesses four putative membrane-interaction elements. Exposing BTV to low pH detaches VP2 and dramatically refolds the dagger and unfurling domains of VP5. Our biochemical and structure-guided-mutagenesis studies support these coordinated pH-sensing mechanisms.
- California NanoSystems Institute, University of California, Los Angeles (UCLA), Los Angeles, California, USA.
Organizational Affiliation: