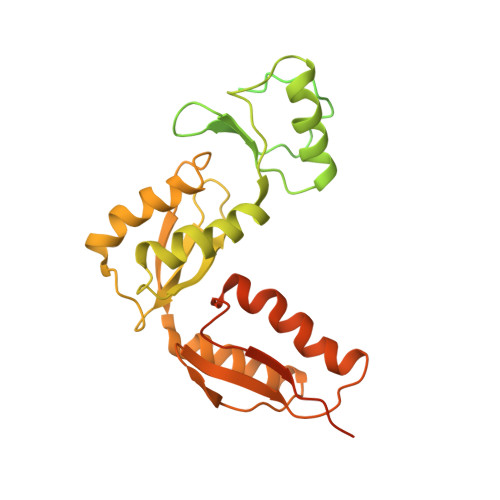
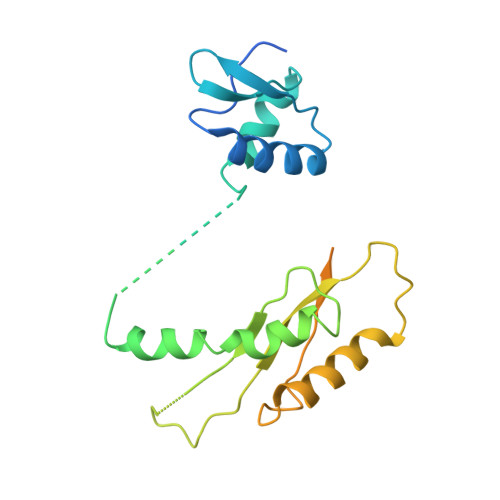
The Modular Structure of the Inner-Membrane Ring Component PrgK Facilitates Assembly of the Type III Secretion System Basal Body.
Bergeron, J.R., Worrall, L.J., De, S., Sgourakis, N.G., Cheung, A.H., Lameignere, E., Okon, M., Wasney, G.A., Baker, D., McIntosh, L.P., Strynadka, N.C.(2015) Structure 23: 161-172
- PubMed: 25533490 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2014.10.021
- Primary Citation Related Structures:
2MKY, 3J6D, 4OYC, 4W4M - PubMed Abstract:
The type III secretion system (T3SS) is a large macromolecular assembly found at the surface of many pathogenic Gram-negative bacteria. Its role is to inject toxic "effector" proteins into the cells of infected organisms. The molecular details of the assembly of this large, multimembrane-spanning complex remain poorly understood. Here, we report structural, biochemical, and functional analyses of PrgK, an inner-membrane component of the prototypical Salmonella typhimurium T3SS. We have obtained the atomic structures of the two ring building globular domains and show that the C-terminal transmembrane helix is not essential for assembly and secretion. We also demonstrate that structural rearrangement of the two PrgK globular domains, driven by an interconnecting linker region, may promote oligomerization into ring structures. Finally, we used electron microscopy-guided symmetry modeling to propose a structural model for the intimately associated PrgH-PrgK ring interaction within the assembled basal body.
- Department of Biochemistry and Molecular Biology, The University of British Columbia, 2350 Health Sciences Mall, Vancouver, BC, V6T 1Z3, Canada; Centre for Blood Research, University of British Columbia, 2350 Health Sciences Mall, Vancouver, BC, V6T 1Z3, Canada.
Organizational Affiliation: