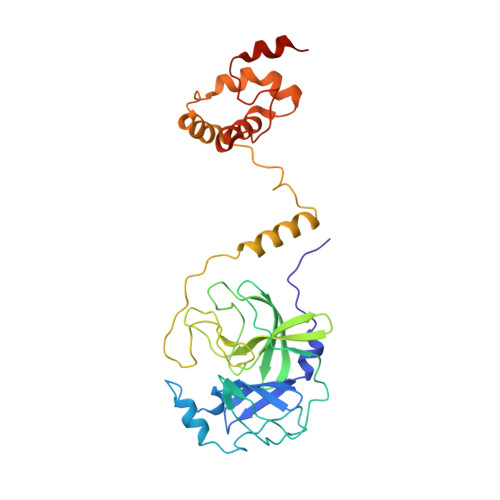
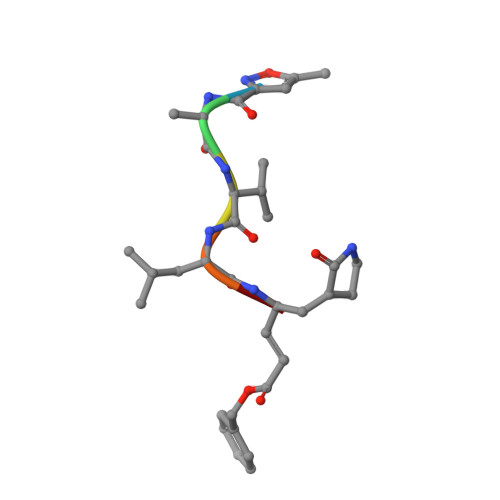
Three-dimensional domain swapping as a mechanism to lock the active conformation in a super-active octamer of SARS-CoV main protease
Zhang, S., Zhong, N., Xue, F., Kang, X., Ren, X., Chen, J., Jin, C., Lou, Z., Xia, B.(2010) Protein Cell 1: 371-383
- PubMed: 21203949 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-010-0044-8
- Primary Citation Related Structures:
3IWM - PubMed Abstract:
Proteolytic processing of viral polyproteins is indispensible for the lifecycle of coronaviruses. The main protease (M(pro)) of SARS-CoV is an attractive target for anti-SARS drug development as it is essential for the polyprotein processing. M(pro) is initially produced as part of viral polyproteins and it is matured by autocleavage. Here, we report that, with the addition of an N-terminal extension peptide, M(pro) can form a domain-swapped dimer. After complete removal of the extension peptide from the dimer, the mature M(pro) self-assembles into a novel super-active octamer (AO-M(pro)). The crystal structure of AO-M(pro) adopts a novel fold with four domain-swapped dimers packing into four active units with nearly identical conformation to that of the previously reported M(pro) active dimer, and 3D domain swapping serves as a mechanism to lock the active conformation due to entanglement of polypeptide chains. Compared with the previously well characterized form of M(pro), in equilibrium between inactive monomer and active dimer, the stable AO-M(pro) exhibits much higher proteolytic activity at low concentration. As all eight active sites are bound with inhibitors, the polyvalent nature of the interaction between AO-M(pro) and its polyprotein substrates with multiple cleavage sites, would make AO-M(pro) functionally much more superior than the M(pro) active dimer for polyprotein processing. Thus, during the initial period of SARS-CoV infection, this novel active form AOM(pro) should play a major role in cleaving polyproteins as the protein level is extremely low. The discovery of AOM(pro) provides new insights about the functional mechanism of M(pro) and its maturation process.
- Beijing Nuclear Magnetic Resonance Center, Peking University, Beijing, 100871, China.
Organizational Affiliation: