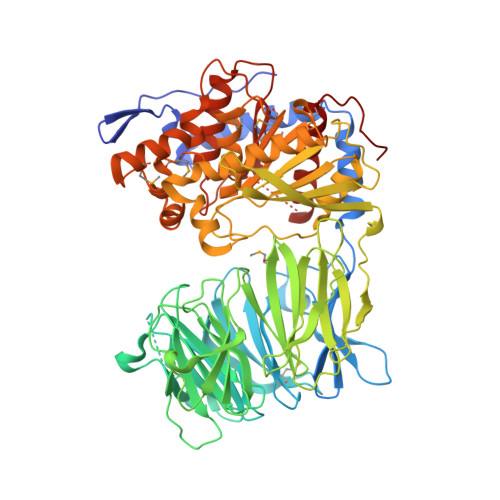
Induced-fit mechanism for prolyl endopeptidase
Li, M., Chen, C., Davies, D.R., Chiu, T.K.(2010) J Biological Chem 285: 21487-21495
- PubMed: 20444688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.092692
- Primary Citation Related Structures:
3IUJ, 3IUL, 3IUM, 3IUN, 3IUQ, 3IUR, 3IVM - PubMed Abstract:
Prolyl peptidases cleave proteins at proline residues and are of importance for cancer, neurological function, and type II diabetes. Prolyl endopeptidase (PEP) cleaves neuropeptides and is a drug target for neuropsychiatric diseases such as post-traumatic stress disorder, depression, and schizophrenia. Previous structural analyses showing little differences between native and substrate-bound structures have suggested a lock-and-key catalytic mechanism. We now directly demonstrate from seven structures of Aeromonus punctata PEP that the mechanism is instead induced fit: the native enzyme exists in a conformationally flexible opened state with a large interdomain opening between the beta-propeller and alpha/beta-hydrolase domains; addition of substrate to preformed native crystals induces a large scale conformational change into a closed state with induced-fit adjustments of the active site, and inhibition of this conformational change prevents substrate binding. Absolute sequence conservation among 28 orthologs of residues at the active site and critical residues at the interdomain interface indicates that this mechanism is conserved in all PEPs. This finding has immediate implications for the use of conformationally targeted drug design to improve specificity of inhibition against this family of proline-specific serine proteases.
- Department of Biochemistry and Molecular Biology, Louisiana State University Health Sciences Center, New Orleans, Louisiana 70112, USA.
Organizational Affiliation: