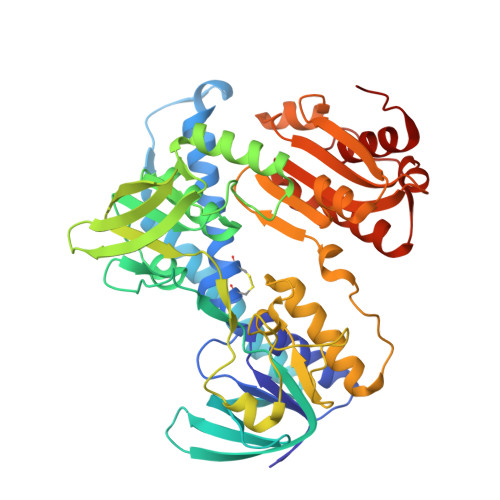
Triazaspirodimethoxybenzoyls as selective inhibitors of mycobacterial lipoamide dehydrogenase .
Bryk, R., Arango, N., Venugopal, A., Warren, J.D., Park, Y.H., Patel, M.S., Lima, C.D., Nathan, C.(2010) Biochemistry 49: 1616-1627
- PubMed: 20078138 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi9016186
- Primary Citation Related Structures:
3II4 - PubMed Abstract:
Mycobacterium tuberculosis (Mtb) remains the leading single cause of death from bacterial infection. Here we explored the possibility of species-selective inhibition of lipoamide dehydrogenase (Lpd), an enzyme central to Mtb's intermediary metabolism and antioxidant defense. High-throughput screening of combinatorial chemical libraries identified triazaspirodimethoxybenzoyls as high-nanomolar inhibitors of Mtb's Lpd that were noncompetitive versus NADH, NAD(+), and lipoamide and >100-fold selective compared to human Lpd. Efficacy required the dimethoxy and dichlorophenyl groups. The structure of an Lpd-inhibitor complex was resolved to 2.42 A by X-ray crystallography, revealing that the inhibitor occupied a pocket adjacent to the Lpd NADH/NAD(+) binding site. The inhibitor did not overlap with the adenosine moiety of NADH/NAD(+) but did overlap with positions predicted to bind the nicotinamide rings in NADH and NAD(+) complexes. The dimethoxy ring occupied a deep pocket adjacent to the FAD flavin ring where it would block coordination of the NADH nicotinamide ring, while the dichlorophenyl group occupied a more exposed pocket predicted to coordinate the NAD(+) nicotinamide. Several residues that are not conserved between the bacterial enzyme and its human homologue were predicted to contribute both to inhibitor binding and to species selectivity, as confirmed for three residues by analysis of the corresponding mutant Mtb Lpd proteins. Thus, nonconservation of residues lining the electron-transfer tunnel in Mtb Lpd can be exploited for development of species-selective Lpd inhibitors.
- Department of Microbiology and Immunology, Weill Cornell Medical College, New York,New York 10065, USA.
Organizational Affiliation: