'Hot' macromolecular crystals.
Koclega, K.D., Chruszcz, M., Zimmerman, M.D., Bujacz, G., Minor, W.(2010) Cryst Growth Des 10: 580-586
- PubMed: 20161694
- DOI: https://doi.org/10.1021/cg900971h
- Primary Citation Related Structures:
3IH2, 3IH3, 3IH4 - PubMed Abstract:
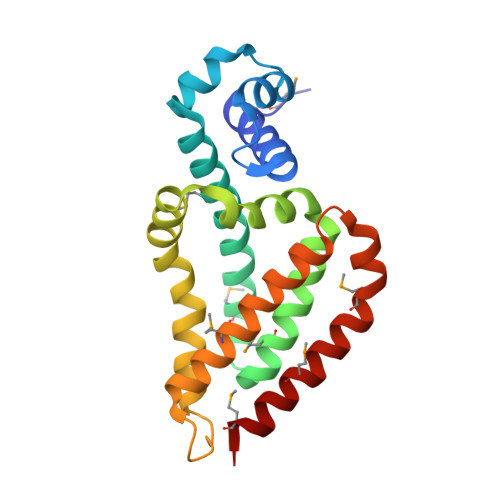
Transcriptional regulator protein TM1030 from the hyperthermophile Thermotoga maritima, as well as its complex with DNA, was crystallized at a wide range of temperatures. Crystallization plates were incubated at 4, 20, 37 and 50° C over 3 weeks. The best crystals of TM1030 in complex with DNA were obtained at 4, 20 and 37° C, while TM1030 alone crystallized almost equally well in all temperatures. The crystals grown at different temperatures were used for X-ray diffraction experiments and their structures were compared. Surprisingly, the models of TM1030 obtained from crystals grown at different temperatures are similar in quality. While there are some examples of structures of proteins grown at elevated temperatures in the PDB, these temperatures appear to be underrepresented. Our studies show that crystals of some proteins may be grown and are stable at broad range of temperatures. We suggest that crystallization experiments at elevated temperatures could be used as a standard part of the crystallization protocol.
- Department of Molecular Physiology and Biological Physics, University of Virginia, 1340 Jefferson Park Avenue, Charlottesville, VA 22908, USA.
Organizational Affiliation: