Identification of 3,4-Dihydroisoquinoline-2(1H)-sulfonamides as Potent Carbonic Anhydrase Inhibitors: Synthesis, Biological Evaluation, and Enzyme-Ligand X-ray Studies.
Gitto, R., Agnello, S., Ferro, S., De Luca, L., Vullo, D., Brynda, J., Mader, P., Supuran, C.T., Chimirri, A.(2010) J Med Chem
- PubMed: 20170095 Search on PubMed
- DOI: https://doi.org/10.1021/jm9014026
- Primary Citation Related Structures:
3IGP - PubMed Abstract:
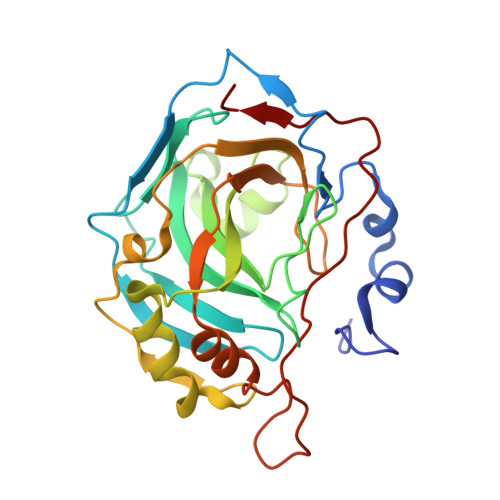
Following previous studies we herein report the exploration of the carbonic anhydrase (CA, EC 4.2.1.1) inhibitory effects and enzyme selectivity of a small class of 1-(cyclo)alkylisoquinolines containing a sulfonamide function considered a key feature for inhibiting CA. The results of enzymatic assays against human (h) CA isoforms, hCA I and hCA II (cytosolic, ubiquitous enzymes), hCA IX (transmembrane, tumor-associated), and hCA XIV (transmembrane), suggested that the presence of C-1 small substituents on isoquinoline scaffold controls both inhibitory potency and selectivity. Some derivatives showed potent hCA IX and hCA XIV inhibitory effects at nanomolar concentrations as well as low affinity for the ubiquitous hCA II. Moreover, we report the X-ray crystal structure of one of these derivatives in complex with dominant human isoform II, thus confirming the sulfonamide--zinc interactions. Finally, the results of docking experiments suggested the hypothetic interactions in the catalytic binding site for the most active and selective hCA IX and hCA XIV inhibitor.
- Dipartimento Farmaco-Chimico, Universita di Messina, Viale Annunziata, I-98168 Messina, Italy. rgitto@pharma.unime.it
Organizational Affiliation: