Identification of orally bioavailable, non-amidine inhibitors of Urokinase Plasminogen Activator (uPA)
West, C.W., Adler, M., Arnaiz, D., Chen, D., Chu, K., Gualtieri, G., Ho, E., Huwe, C., Light, D., Phillips, G., Pulk, R., Sukovich, D., Whitlow, M., Yuan, S., Bryant, J.(2009) Bioorg Med Chem Lett 19: 5712-5715
- PubMed: 19703768 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2009.08.008
- Primary Citation Related Structures:
3IG6 - PubMed Abstract:
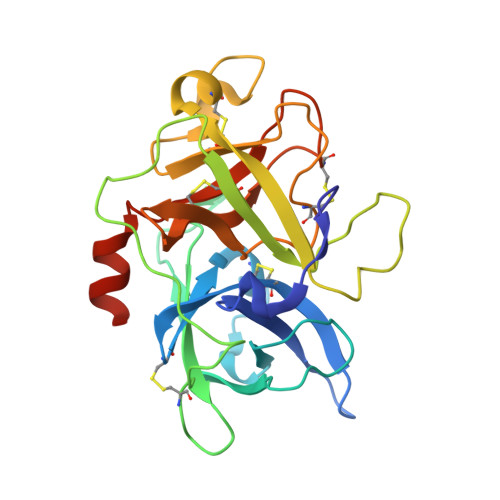
In this Letter we report the synthesis and evaluation of a series of non-amidine inhibitors of Urokinase Plasminogen Activator (uPA). Starting from compound 1, a significant change provided compounds in which the amidine, binding in the S1 pocket, was replaced with a primary amine. Further modifications led to the identification of potent, selective, and orally bioavailable uPA inhibitors.
- Department of Medicinal Chemistry, Berlex Biosciences, 2600 Hilltop Drive, Richmond, CA 94804, United States. cwest@icagen.com
Organizational Affiliation: