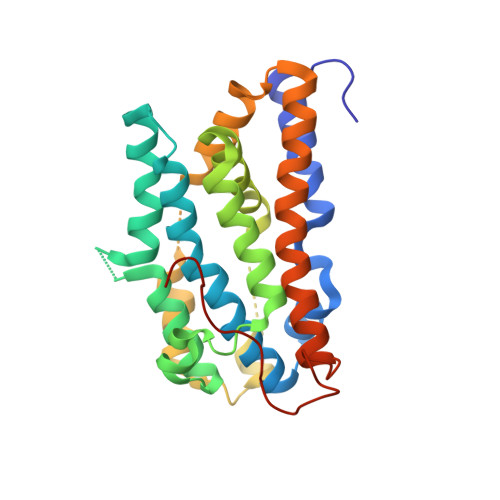
Structural studies of mutant forms of the PQQ-forming enzyme PqqC in the presence of product and substrate
Puehringer, S., RoseFigura, J., Metlitzky, M., Toyama, H., Klinman, J.P., Schwarzenbacher, R.(2010) Proteins 78: 2554-2562
- PubMed: 20602352 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22769
- Primary Citation Related Structures:
3HLX, 3HML, 3HNH - PubMed Abstract:
Pyrroloquinoline quinone [4,5-dihydro-4,5-dioxo-1H-pyrrolo[2,3-f]quinoline-2,7,9-tricarboxylic acid (PQQ)] is a bacterial cofactor in numerous alcohol dehydrogenases including methanol dehydrogenase and glucose dehydrogenase. Its biosynthesis in Klebsiella pneumoniae is facilitated by six genes, pqqABCDEF and proceeds by an unknown pathway. PqqC is one of two metal free oxidases of known structure and catalyzes the last step of PQQ biogenesis which involves a ring closure and an eight-electron oxidation of the substrate [3a-(2-amino-2-carboxyethyl)-4,5-dioxo-4,5,6,7,8,9-hexahydroquinoline-7,9-dicarboxylic acid (AHQQ)]. PqqC has 14 conserved active site residues, which have previously been shown to be in close contact with bound PQQ. Herein, we describe the structures of three PqqC active site variants, H154S, Y175F, and the double mutant R179S/Y175S. The H154S crystal structure shows that, even with PQQ bound, the enzyme is still in the "open" conformation with helices alpha5b and alpha6 unfolded and the active site solvent accessible. The Y175F PQQ complex crystal structure reveals the closed conformation indicating that Y175 is not required for the conformational change. The R179S/Y175S AHQQ complex crystal structure is the most mechanistically informative, indicating an open conformation with a reaction intermediate trapped in the active site. The intermediate seen in R179S/Y175S is tricyclic but nonplanar, implying that it has not undergone oxidation. These studies implicate a stepwise process in which substrate binding leads to the generation of the closed protein conformation, with the latter playing a critical role in O(2) binding and catalysis.
- Department of Molecular Biology, University of Salzburg, 5020 Salzburg, Austria.
Organizational Affiliation: