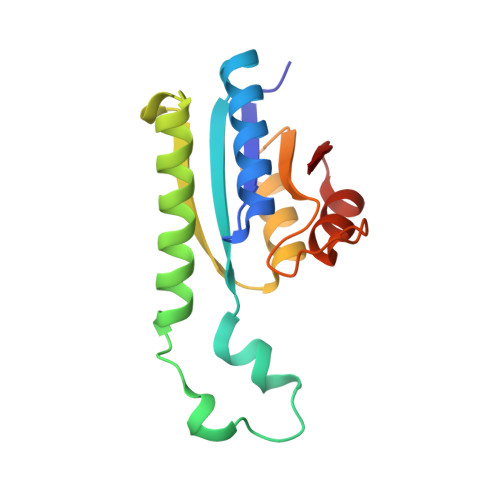
Structure and function of the universal stress protein TeaD and its role in regulating the ectoine transporter TeaABC of Halomonas elongata DSM 2581(T)
Schweikhard, E.S., Kuhlmann, S.I., Kunte, H.J., Grammann, K., Ziegler, C.M.(2010) Biochemistry 49: 2194-2204
- PubMed: 20113006 Search on PubMed
- DOI: https://doi.org/10.1021/bi9017522
- Primary Citation Related Structures:
3HGM - PubMed Abstract:
The halophilic bacterium Halomonas elongata takes up the compatible solute ectoine via the osmoregulated TRAP transporter TeaABC. A fourth orf (teaD) is located adjacent to the teaABC locus that encodes a putative universal stress protein (USP). By RT-PCR experiments we proved a cotranscription of teaD along with teaABC. Deletion of teaD resulted in an enhanced uptake for ectoine by the transporter TeaABC and hence a negative activity regulation of TeaABC by TeaD. A transcriptional regulation via DNA binding could be excluded. ATP binding to native TeaD was shown by HPLC, and the crystal structure of TeaD was solved in complex with ATP to a resolution of 1.9 A by molecular replacement. TeaD forms a dimer-dimer complex with one ATP molecule bound to each monomer, which has a Rossmann-like alpha/beta overall fold. Our results reveal an ATP-dependent oligomerization of TeaD, which might have a functional role in the regulatory mechanism of TeaD. USP-encoding orfs, which are located adjacent to genes encoding for TeaABC homologues, could be identified in several other organisms, and their physiological role in balancing the internal cellular ectoine pool is discussed.
- Department of Structural Biology, Max Planck Institute of Biophysics, Max-von-Laue-Strasse 3, 60438 Frankfurt am Main, Germany.
Organizational Affiliation: