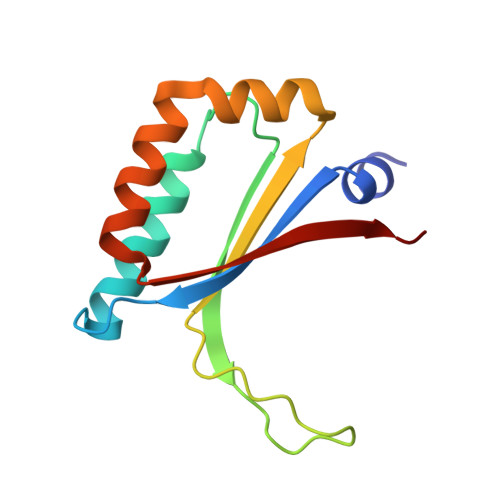
Crystal structure and catalytic mechanism of 4-methylmuconolactone methylisomerase
Marin, M., Heinz, D.W., Pieper, D.H., Klink, B.U.(2009) J Biological Chem 284: 32709-32716
- PubMed: 19801657 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.024604
- Primary Citation Related Structures:
3HDS, 3HF5, 3HFK - PubMed Abstract:
When methyl-substituted aromatic compounds are degraded via ortho (intradiol)-cleavage of 4-methylcatechol, the dead-end metabolite 4-methylmuconolactone (4-ML) is formed. Degradation of 4-ML has only been described in few bacterial species, including Pseudomonas reinekei MT1. The isomerization of 4-ML to 3-methylmuconolactone (3-ML) is the first step required for the mineralization of 4-ML and is catalyzed by an enzyme termed 4-methylmuconolactone methylisomerase (MLMI). We identified the gene encoding MLMI in P. reinekei MT1 and solved the crystal structures of MLMI in complex with 3-ML at 1.4-A resolution, with 4-ML at 1.9-A resolution and with a MES buffer molecule at 1.45-A resolution. MLMI exhibits a ferredoxin-like fold and assembles as a tight functional homodimeric complex. We were able to assign the active site clefts of MLMI from P. reinekei MT1 and of the homologous MLMI from Cupriavidus necator JMP134, which has previously been crystallized in a structural genomics project. Kinetic and structural analysis of wild-type MLMI and variants created by site-directed mutagenesis indicate Tyr-39 and His-26 to be the most probable catalytic residues. The previously proposed involvement of Cys-67 in covalent catalysis can now be excluded. Residue His-52 was found to be important for substrate affinity, with only marginal effect on catalytic activity. Based on these results, a novel catalytic mechanism for the isomerization of 4-ML to 3-ML by MLMI, involving a bislactonic intermediate, is proposed. This broadens the knowledge about the diverse group of proteins exhibiting a ferredoxin-like fold.
- Department of Microbial Pathogenesis, Helmholtz Centre for Infection Research, Inhoffenstrasse 7, 38124 Braunschweig, Germany.
Organizational Affiliation: