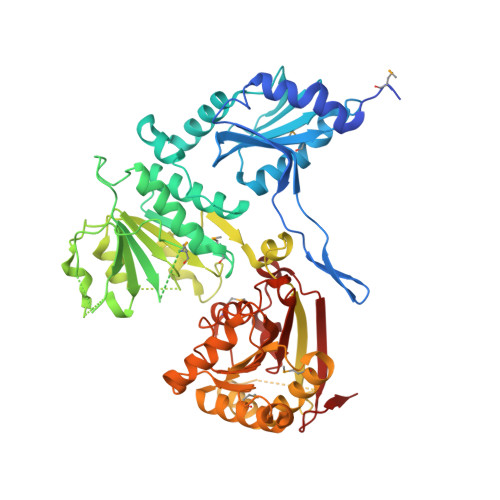
The fragment structure of a putative HsdR subunit of a type I restriction enzyme from Vibrio vulnificus YJ016: implications for DNA restriction and translocation activity
Uyen, N.T., Park, S.Y., Choi, J.W., Lee, H.J., Nishi, K., Kim, J.S.(2009) Nucleic Acids Res
- PubMed: 19625490 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkp603
- Primary Citation Related Structures:
3H1T - PubMed Abstract:
Among four types of bacterial restriction enzymes that cleave a foreign DNA depending on its methylation status, type I enzymes composed of three subunits are interesting because of their unique DNA cleavage and translocation mechanisms performed by the restriction subunit (HsdR). The elucidated N-terminal fragment structure of a putative HsdR subunit from Vibrio vulnificus YJ016 reveals three globular domains. The nucleolytic core within an N-terminal nuclease domain (NTD) is composed of one basic and three acidic residues, which include a metal-binding site. An ATP hydrolase (ATPase) site at the interface of two RecA-like domains (RDs) is located close to the probable DNA-binding site for translocation, which is far from the NTD nucleolytic core. Comparison of relative domain arrangements with other functionally related ATP and/or DNA complex structures suggests a possible translocation and restriction mechanism of the HsdR subunit. Furthermore, careful analysis of its sequence and structure implies that a linker helix connecting two RDs and an extended region within the nuclease domain may play a central role in switching the DNA translocation into the restriction activity.
- Interdisciplinary Graduate Program in Molecular Medicine, Gwangju 501-746, Korea.
Organizational Affiliation: