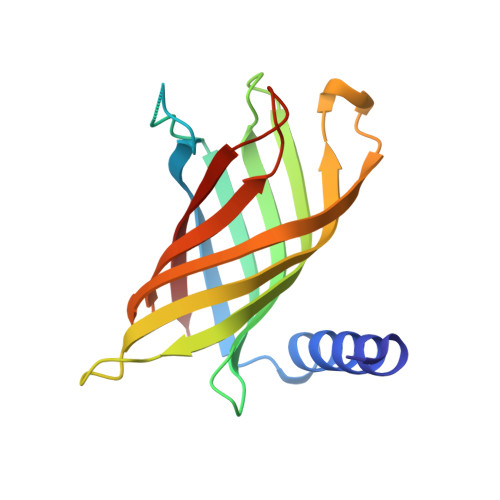
PagP crystallized from SDS/cosolvent reveals the route for phospholipid access to the hydrocarbon ruler.
Cuesta-Seijo, J.A., Neale, C., Khan, M.A., Moktar, J., Tran, C.D., Bishop, R.E., Pomes, R., Prive, G.G.(2010) Structure 18: 1210-1219
- PubMed: 20826347 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2010.06.014
- Primary Citation Related Structures:
3GP6 - PubMed Abstract:
Enzymatic reactions involving bilayer lipids occur in an environment with strict physical and topological constraints. The integral membrane enzyme PagP transfers a palmitoyl group from a phospholipid to lipid A in order to assist Escherichia coli in evading host immune defenses during infection. PagP measures the palmitoyl group with an internal hydrocarbon ruler that is formed in the interior of the eight-stranded antiparallel β barrel. The access and egress of the palmitoyl group is thought to take a lateral route from the bilayer phase to the barrel interior. Molecular dynamics, mutagenesis, and a 1.4 A crystal structure of PagP in an SDS / 2-methyl-2,4-pentanediol (MPD) cosolvent system reveal that phospholipid access occurs at the crenel present between strands F and G of PagP. In this way, the phospholipid head group can remain exposed to the cell exterior while the lipid acyl chain remains in a predominantly hydrophobic environment as it translocates to the protein interior.
- Division of Cancer Genomics and Proteomics, Ontario Cancer Institute and Campbell Family Cancer Research Institute, 101 College Street, Toronto, ON M5G 1L7, Canada.
Organizational Affiliation: