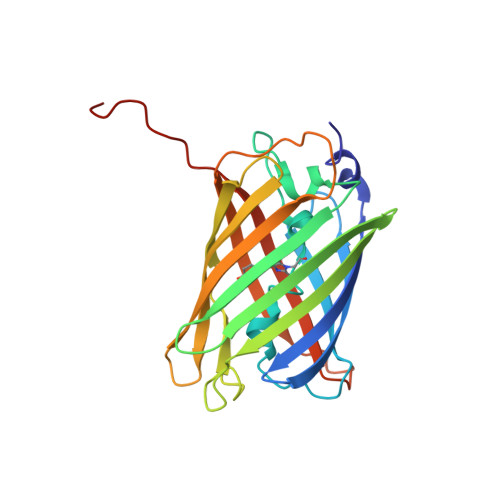
Structural basis for phototoxicity of the genetically encoded photosensitizer KillerRed.
Pletnev, S., Gurskaya, N.G., Pletneva, N.V., Lukyanov, K.A., Chudakov, D.M., Martynov, V.I., Popov, V.O., Kovalchuk, M.V., Wlodawer, A., Dauter, Z., Pletnev, V.(2009) J Biological Chem 284: 32028-32039
- PubMed: 19737938 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.054973
- Primary Citation Related Structures:
3GB3, 3GL4 - PubMed Abstract:
KillerRed is the only known fluorescent protein that demonstrates notable phototoxicity, exceeding that of the other green and red fluorescent proteins by at least 1,000-fold. KillerRed could serve as an instrument to inactivate target proteins or to kill cell populations in photodynamic therapy. However, the nature of KillerRed phototoxicity has remained unclear, impeding the development of more phototoxic variants. Here we present the results of a high resolution crystallographic study of KillerRed in the active fluorescent and in the photobleached non-fluorescent states. A unique and striking feature of the structure is a water-filled channel reaching the chromophore area from the end cap of the beta-barrel that is probably one of the key structural features responsible for phototoxicity. A study of the structure-function relationship of KillerRed, supported by structure-based, site-directed mutagenesis, has also revealed the key residues most likely responsible for the phototoxic effect. In particular, Glu(68) and Ser(119), located adjacent to the chromophore, have been assigned as the primary trigger of the reaction chain.
- Synchrotron Radiation Research Section, Macromolecular Crystallography Laboratory, National Cancer Institute/SAIC-Frederick Inc., Frederick, Maryland 21702, USA.
Organizational Affiliation: