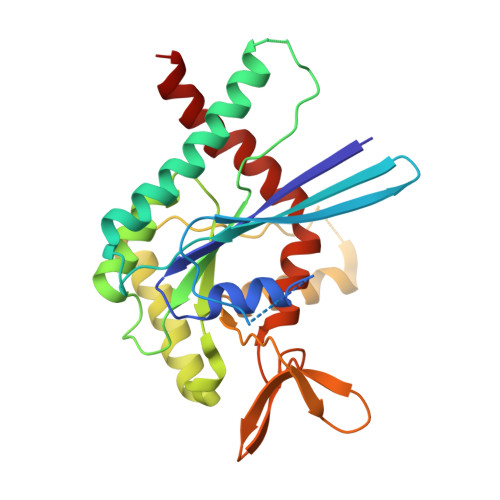
GTP-induced conformational changes in septins and implications for function
Sirajuddin, M., Farkasovsky, M., Zent, E., Wittinghofer, A.(2009) Proc Natl Acad Sci U S A 106: 16592-16597
- PubMed: 19805342 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0902858106
- Primary Citation Related Structures:
3FTQ - PubMed Abstract:
Septins constitute a group of GTP-binding proteins involved in cytokinesis and other essential cellular functions. They form heterooligomeric complexes that polymerize into nonpolar filaments and are dynamic during different stages of the cell cycle. Posttranslational modifications and interacting partners are widely accepted regulators of septin filament function, but the contribution of nucleotide is undefined due to a lack of detailed structural information. Previous low-resolution structures showed that the G domain assembles into a linear polymer with 2 different interfaces involving the N and C termini and the G binding sites. Here we report the crystal structure of SEPT2 bound to GppNHp at 2.9 A resolution. GTP binding induces conformational changes in the switch regions at the G interfaces, which are transmitted to the N-terminal helix and also affect the NC interface. Biochemical studies and sequence alignment suggest that a threonine, which is conserved in certain subgroups of septins, is responsible for GTP hydrolysis. Although this threonine is not present in yeast CDC3 and CDC11, its mutation in CDC10 and CDC12 induces temperature sensitivity. Highly conserved contact residues identified in the G interface are shown to be necessary for Cdc3-10, but not Cdc11-12, heterodimer formation and cell growth in yeast. Based on our findings, we propose that GTP binding/hydrolysis and the nature of the nucleotide influence the stability of interfaces in heterooligomeric and polymeric septins and are required for proper septin filament assembly/disassembly. These data also offer a first rationale for subdividing human septins into different functional subgroups.
- Abteilung Strukturelle Biologie, Max-Planck-Institut für molekulare Physiologie, Dortmund, Germany.
Organizational Affiliation: