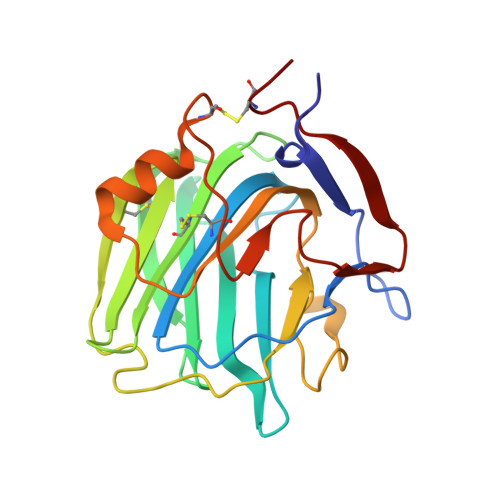
Crystal structures of Limulus SAP-like pentraxin reveal two molecular aggregations.
Shrive, A.K., Burns, I., Chou, H.T., Stahlberg, H., Armstrong, P.B., Greenhough, T.J.(2009) J Mol Biology 386: 1240-1254
- PubMed: 19452596 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.01.008
- Primary Citation Related Structures:
3FLP, 3FLR, 3FLT - PubMed Abstract:
The serum-amyloid-P-component-like pentraxin from Limulus polyphemus, a recently discovered pentraxin species and important effector protein of the hemolymph immune system, displays two distinct doubly stacked cyclic molecular aggregations, heptameric and octameric. The refined three-dimensional structures determined by X-ray crystallography, both based on the same cDNA sequence, show that each aggregate is constructed from a similar dimer of protomers, which is repeated to make up the ring structure. The native octameric form has been refined at a resolution of 3 A, the native heptameric form at 2.3 A, and the phosphoethanolamine (PE)-bound octameric form at 2.7 A. The existence of the hitherto undescribed heptameric form was confirmed by single-particle analysis using cryo-electron microscopy. In the native structures, the calcium-binding site is similar to that in human pentraxins, with two calcium ions bound in each subunit. Upon binding PE, however, each subunit binds a third calcium ion, with all three calcium ions contributing to the binding and orientation of the bound phosphate group within the ligand-binding pocket. While the phosphate is well-defined in the electron density, the ethanolamine group is poorly defined, suggesting structural and binding variabilities of this group. Although sequence homology with human serum amyloid P component is relatively low, structural homology is high, with very similar overall folds and a common affinity for PE. This is due, in part, to a "topological" equivalence of side-chain position. Identical side chains that are important in both function and fold, from different regions of the sequence in human and Limulus structures, occupy similar space within the overall subunit fold. Sequence and structure alignment, based on the refined three-dimensional structures presented here and the known horseshoe crab pentraxin sequences, suggest that adaptation and refinement of C-reactive-protein-mediated immune responses in these ancient creatures lacking antibody-based immunity are based on adaptation by gene duplication.
- Institute of Science and Technology in Medicine, School of Life Sciences, Keele University, Staffordshire ST5 5BG, UK.
Organizational Affiliation: