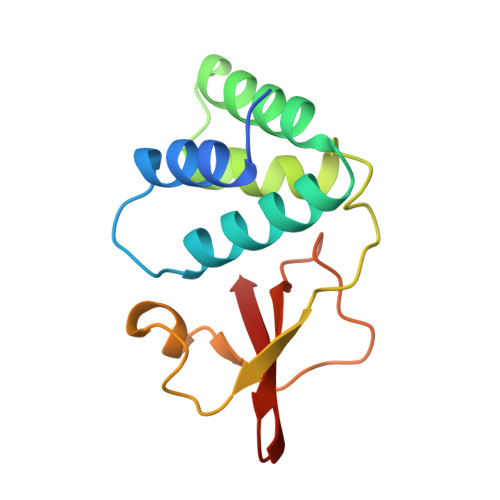
Structure of the Ebola VP35 interferon inhibitory domain.
Leung, D.W., Ginder, N.D., Fulton, D.B., Nix, J., Basler, C.F., Honzatko, R.B., Amarasinghe, G.K.(2009) Proc Natl Acad Sci U S A 106: 411-416
- PubMed: 19122151 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0807854106
- Primary Citation Related Structures:
3FKE - PubMed Abstract:
Ebola viruses (EBOVs) cause rare but highly fatal outbreaks of viral hemorrhagic fever in humans, and approved treatments for these infections are currently lacking. The Ebola VP35 protein is multifunctional, acting as a component of the viral RNA polymerase complex, a viral assembly factor, and an inhibitor of host interferon (IFN) production. Mutation of select basic residues within the C-terminal half of VP35 abrogates its dsRNA-binding activity, impairs VP35-mediated IFN antagonism, and attenuates EBOV growth in vitro and in vivo. Because VP35 contributes to viral escape from host innate immunity and is required for EBOV virulence, understanding the structural basis for VP35 dsRNA binding, which correlates with suppression of IFN activity, is of high importance. Here, we report the structure of the C-terminal VP35 IFN inhibitory domain (IID) solved to a resolution of 1.4 A and show that VP35 IID forms a unique fold. In the structure, we identify 2 basic residue clusters, one of which is important for dsRNA binding. The dsRNA binding cluster is centered on Arg-312, a highly conserved residue required for IFN inhibition. Mutation of residues within this cluster significantly changes the surface electrostatic potential and diminishes dsRNA binding activity. The high-resolution structure and the identification of the conserved dsRNA binding residue cluster provide opportunities for antiviral therapeutic design. Our results suggest a structure-based model for dsRNA-mediated innate immune antagonism by Ebola VP35 and other similarly constructed viral antagonists.
- Department of Biochemistry, Biophysics and Molecular Biology, Iowa State University, Ames, IA 50011, USA.
Organizational Affiliation: