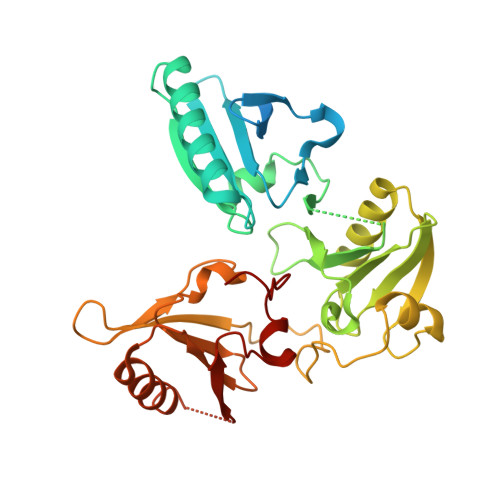
The crystal structure of the C-terminus of adseverin reveals the actin-binding interface.
Chumnarnsilpa, S., Lee, W.L., Nag, S., Kannan, B., Larsson, M., Burtnick, L.D., Robinson, R.C.(2009) Proc Natl Acad Sci U S A 106: 13719-13724
- PubMed: 19666531 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0812383106
- Primary Citation Related Structures:
3FG6 - PubMed Abstract:
Adseverin is a member of the calcium-regulated gelsolin superfamily of actin severing and capping proteins. Adseverin comprises 6 homologous domains (A1-A6), which share 60% identity with the 6 domains from gelsolin (G1-G6). Adseverin is truncated in comparison to gelsolin, lacking the C-terminal extension that masks the F-actin binding site in calcium-free gelsolin. Biochemical assays have indicated differences in the interaction of the C-terminal halves of adseverin and gelsolin with actin. Gelsolin contacts actin through a major site on G4 and a minor site on G6, whereas adseverin uses a site on A5. Here, we present the X-ray structure of the activated C-terminal half of adseverin (A4-A6). This structure is highly similar to that of the activated form of the C-terminal half of gelsolin (G4-G6), both in arrangement of domains and in the 3 bound calcium ions. Comparative analysis of the actin-binding surfaces observed in the G4-G6/actin structure suggests that adseverin in this conformation will also be able to interact with actin through A4 and A6, whereas the A5 surface is obscured. A single residue mutation in A4-A6 located at the predicted A4/actin interface completely abrogates actin sequestration. A model of calcium-free adseverin, constructed from the structure of gelsolin, predicts that in the absence of a gelsolin-like C-terminal extension the interaction between A2 and A6 provides the steric inhibition to prevent interaction with F-actin. We propose that calcium binding to the N terminus of adseverin dominates the activation process to expose the F-actin binding site on A2.
- Institute of Molecular and Cell Biology, Agency for Science, Technology, and Research (ASTAR), 61 Biopolis Drive, Proteos, Singapore 138673.
Organizational Affiliation: