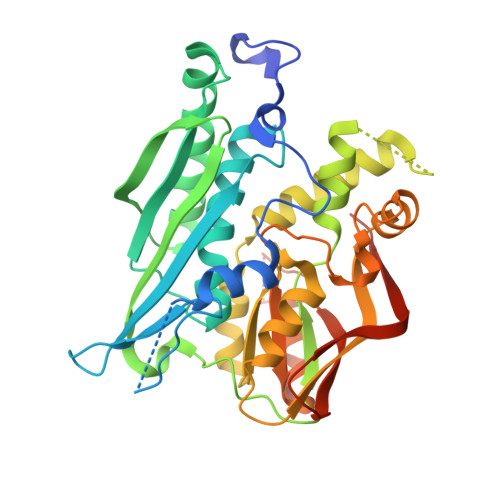
Crystal structures of catalytic intermediates of human selenophosphate synthetase 1.
Wang, K.T., Wang, J., Li, L.F., Su, X.D.(2009) J Mol Biology 390: 747-759
- PubMed: 19477186
- DOI: https://doi.org/10.1016/j.jmb.2009.05.032
- Primary Citation Related Structures:
3FD5, 3FD6 - PubMed Abstract:
Selenophosphate synthetase catalyzes the synthesis of the highly active selenium donor molecule selenophosphate, a key intermediate in selenium metabolism. We have determined the high-resolution crystal structure of human selenophosphate synthetase 1 (hSPS1). An unexpected reaction intermediate, with a tightly bound phosphate and ADP at the active site has been captured in the structure. An enzymatic assay revealed that hSPS1 possesses low ADP hydrolysis activity in the presence of phosphate. Our structural and enzymatic results suggest that consuming the second high-energy phosphoester bond of ATP could protect the labile product selenophosphate during catalytic reaction. We solved another hSPS1 structure with potassium ions at the active sites. Comparing the two structures, we were able to define the monovalent cation-binding site of the enzyme. The detailed mechanism of the ADP hydrolysis step and the exact function of the monovalent cation for hSPS1 catalytic reaction are proposed.
- National Laboratory of Protein Engineering and Plant Genetic Engineering, College of Life Sciences, Peking University, Beijing, P.R. China.
Organizational Affiliation: