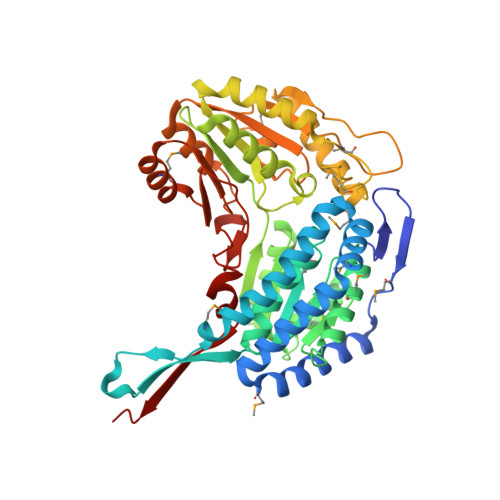
Structure and activity of the NAD(P)(+) -dependent succinate semialdehyde dehydrogenase YneI from Salmonella typhimurium.
Zheng, H., Beliavsky, A., Tchigvintsev, A., Brunzelle, J.S., Brown, G., Flick, R., Evdokimova, E., Wawrzak, Z., Mahadevan, R., Anderson, W.F., Savchenko, A., Yakunin, A.F.(2013) Proteins 81: 1031-1041
- PubMed: 23229889
- DOI: https://doi.org/10.1002/prot.24227
- Primary Citation Related Structures:
3EFV, 3ETF - PubMed Abstract:
Aldehyde dehydrogenases are found in all organisms and play an important role in the metabolic conversion and detoxification of endogenous and exogenous aldehydes. Genomes of many organisms including Escherichia coli and Salmonella typhimurium encode two succinate semialdehyde dehydrogenases with low sequence similarity and different cofactor preference (YneI and GabD). Here, we present the crystal structure and biochemical characterization of the NAD(P)(+)-dependent succinate semialdehyde dehydrogenase YneI from S. typhimurium. This enzyme shows high activity and affinity toward succinate semialdehyde and exhibits substrate inhibition at concentrations of SSA higher than 0.1 mM. YneI can use both NAD(+) and NADP(+) as cofactors, although affinity to NAD(+) is 10 times higher. High resolution crystal structures of YneI were solved in a free state (1.85 Å) and in complex with NAD(+) (1.90 Å) revealing a two domain protein with the active site located in the interdomain interface. The NAD(+) molecule is bound in the long channel with its nicotinamide ring positioned close to the side chain of the catalytic Cys268. Site-directed mutagenesis demonstrated that this residue, as well as the conserved Trp136, Glu365, and Asp426 are important for activity of YneI, and that the conserved Lys160 contributes to the enzyme preference to NAD(+) . Our work has provided further insight into the molecular mechanisms of substrate selectivity and activity of succinate semialdehyde dehydrogenases.
- Department of Chemical Engineering and Applied Chemistry, Banting and Best Department of Medical Research, University of Toronto, Toronto, Ontario, Canada.
Organizational Affiliation: