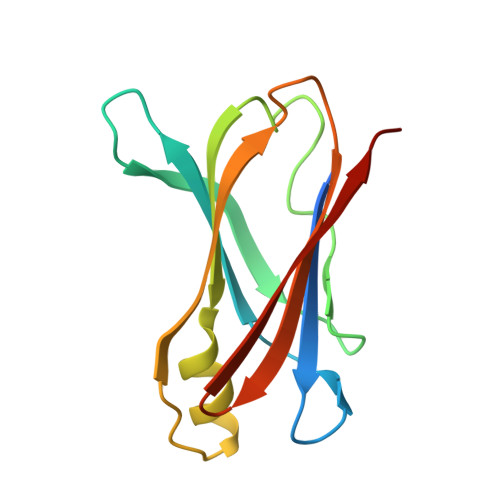
Toward optimization of the second aryl substructure common to transthyretin amyloidogenesis inhibitors using biochemical and structural studies.
Johnson, S.M., Connelly, S., Wilson, I.A., Kelly, J.W.(2009) J Med Chem 52: 1115-1125
- PubMed: 19191553 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm801347s
- Primary Citation Related Structures:
3ESN, 3ESO, 3ESP - PubMed Abstract:
Transthyretin (TTR) amyloidogenesis inhibitors are typically composed of two aromatic rings and a linker. We have previously established optimal structures for one aromatic ring and the linker. Herein, we employ a suboptimal linker and an optimal aryl-X substructure to rank order the desirability of aryl-Z substructures--using a library of 56 N-(3,5-dibromo-4-hydroxyphenyl)benzamides. Coconsideration of amyloid inhibition potency and ex vivo plasma TTR binding selectivity data reveal that 2,6, 2,5, 2, 3,4,5, and 3,5 substituted aryls bearing small substituents generate the most potent and selective inhibitors, in descending order. These benzamides generally lack undesirable thyroid hormone receptor binding and COX-1 inhibition activity. Three high-resolution TTR.inhibitor crystal structures (1.31-1.35 A) provide insight into why these inhibitors are potent and selective, enabling future structure-based design of TTR kinetic stabilizers.
- Department of Chemistry, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, California 92037, USA.
Organizational Affiliation: