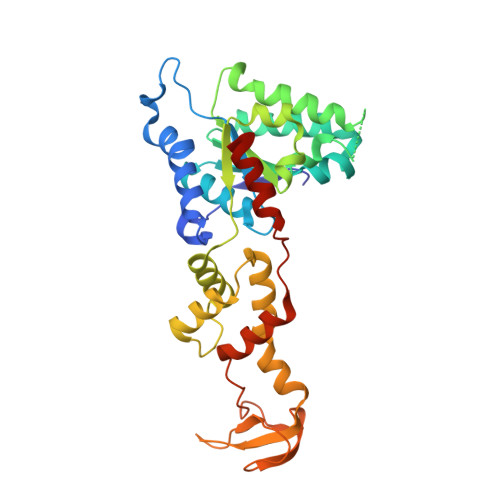
Biochemical and structural studies of yeast vps4 oligomerization.
Gonciarz, M.D., Whitby, F.G., Eckert, D.M., Kieffer, C., Heroux, A., Sundquist, W.I., Hill, C.P.(2008) J Mol Biology 384: 878-895
- PubMed: 18929572
- DOI: https://doi.org/10.1016/j.jmb.2008.09.066
- Primary Citation of Related Structures:
3EIE, 3EIH - PubMed Abstract:
The ESCRT (endosomal sorting complexes required for transport) pathway functions in vesicle formation at the multivesicular body, the budding of enveloped RNA viruses such as HIV-1, and the final abscission stage of cytokinesis. As the only known enzyme in the ESCRT pathway, the AAA ATPase (ATPase associated with diverse cellular activities) Vps4 provides the energy required for multiple rounds of vesicle formation. Like other Vps4 proteins, yeast Vps4 cycles through two states: a catalytically inactive disassembled state that we show here is a dimer and a catalytically active higher-order assembly that we have modeled as a dodecamer composed of two stacked hexameric rings. We also report crystal structures of yeast Vps4 proteins in the apo- and ATPgammaS [adenosine 5'-O-(3-thiotriphosphate)]-bound states. In both cases, Vps4 subunits assembled into continuous helices with 6-fold screw axes that are analogous to helices seen previously in other Vps4 crystal forms. The helices are stabilized by extensive interactions between the large and small AAA ATPase domains of adjacent Vps4 subunits, suggesting that these contact surfaces may be used to build both the catalytically active dodecamer and catalytically inactive dimer. Consistent with this model, we have identified interface mutants that specifically inhibit Vps4 dimerization, dodecamerization, or both. Thus, the Vps4 dimer and dodecamer likely form distinct but overlapping interfaces. Finally, our structural studies have allowed us to model the conformation of a conserved loop (pore loop 2) that is predicted to form an arginine-rich pore at the center of one of the Vps4 hexameric rings. Our mutational analyses demonstrate that pore loop 2 residues Arg241 and Arg251 are required for efficient HIV-1 budding, thereby supporting a role for this "arginine collar" in Vps4 function.
- Department of Biochemistry, University of Utah, Salt Lake City, UT 84112-5650, USA.
Organizational Affiliation: