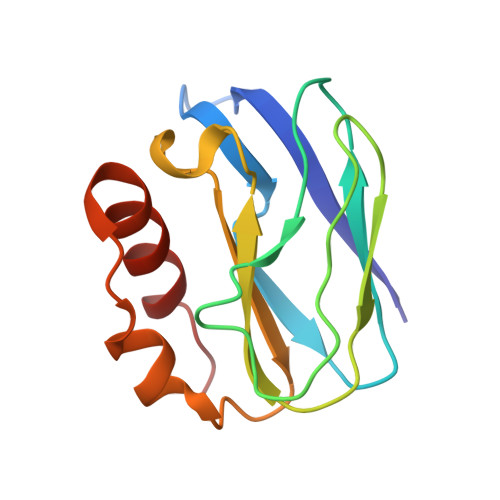
Atomic resolution structure of pseudoazurin from the methylotrophic denitrifying bacterium Hyphomicrobium denitrificans: structural insights into its spectroscopic properties
Hira, D., Nojiri, M., Suzuki, S.(2009) Acta Crystallogr D Biol Crystallogr 65: 85-92
- PubMed: 19153470
- DOI: https://doi.org/10.1107/S0907444908040195
- Primary Citation Related Structures:
3EF4 - PubMed Abstract:
The crystal structure of native pseudoazurin (HdPAz) from the methylotrophic denitrifying bacterium Hyphomicrobium denitrificans has been determined at a resolution of 1.18 A. After refinement with SHELX employing anisotropic displacement parameters and riding H atoms, R(work) and R(free) were 0.135 and 0.169, respectively. Visualization of the anisotropic displacement parameters as thermal ellipsoids provided insight into the atomic motion within the perturbed type 1 Cu site. The asymmetric unit includes three HdPAz molecules which are tightly packed by head-to-head cupredoxin dimer formation. The shape of the Cu-atom ellipsoid implies significant vibrational motion diagonal to the equatorial xy plane defined by the three ligands (two His and one Cys). The geometric parameters of the type 1 Cu site in the HdPAz structure differ unambiguously from those of other pseudoazurins. It is demonstrated that their structural aspects are consistent with the unique visible absorption spectrum.
- Bioinorganic Chemistry Laboratory, Department of Chemistry, Graduate School of Science, Osaka University, 1-1 Machikaneyama, Toyonaka, Osaka 560-0043, Japan.
Organizational Affiliation: