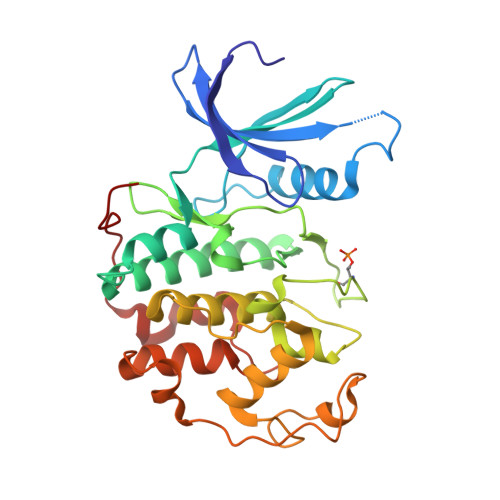
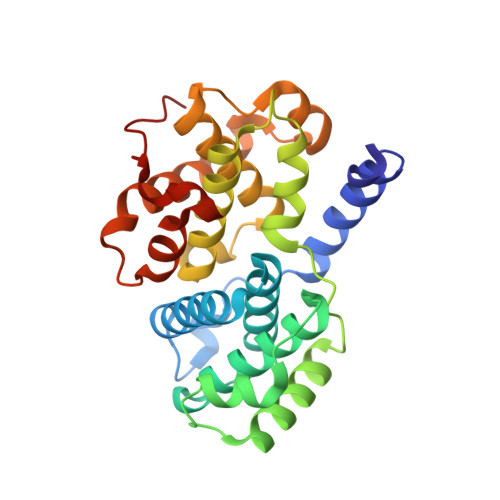
N-&-N, a new class of cell death-inducing kinase inhibitors derived from the purine roscovitine.
Bettayeb, K., Sallam, H., Ferandin, Y., Popowycz, F., Fournet, G., Hassan, M., Echalier, A., Bernard, P., Endicott, J., Joseph, B., Meijer, L.(2008) Mol Cancer Ther 7: 2713-2724
- PubMed: 18790752 Search on PubMed
- DOI: https://doi.org/10.1158/1535-7163.MCT-08-0080
- Primary Citation Related Structures:
3DOG - PubMed Abstract:
Cyclin-dependent kinases (CDKs) and their regulators show frequent abnormalities in tumors. Ten low molecular weight pharmacologic inhibitors of CDKs are currently in clinical trials against various cancers, including the 2,6,9-trisubstituted purine (R)-roscovitine (CYC202/Seliciclib). We here report the characterization of N-&-N1, a bioisoster of roscovitine displaying improved antitumoral properties. N-&-N1 shows exquisite selectivity for CDKs, with 2- to 3-fold enhanced potency compared with (R)-roscovitine. Inhibition of retinoblastoma protein phosphorylation and RNA polymerase II Ser2 phosphorylation in neuroblastoma SH-SY5Y cells exposed to N-&-N1 indicates that N-&-N1 is able to inhibit CDKs in a cellular context. N-&-N1 also down-regulates the expression of RNA polymerase. Cocrystal structures of N-&-N1 and (R)-roscovitine in complex with CDK2/cyclin A reveal that both inhibitors adopt similar binding modes. A competitive assay shows that, compared with (R)-roscovitine, N-&-N1 has reduced affinity for Erk2 and pyridoxal kinase. N-&-N1 triggers cell death in a panel of diverse cell lines. Cell death is accompanied by events characteristic of apoptosis: cytochrome c release, activation of effector caspases, and poly(ADP-ribose) polymerase cleavage. Induction of p53 and p21CIP1 and down-regulation of the Mcl-1 antiapoptotic factor were also observed. Studies in mice show that N-&-N1 has pharmacokinetics properties similar to those of (R)-roscovitine. Altogether, these results show that analogues of (R)-roscovitine can be designed with improved antitumor potential.
- Centre National de la Recherche Scientifique, Protein Phosphorylation and Human Disease Group, UPS2682, Station Biologique, B.P. 74, 29682 Roscoff cedex, Bretagne, France.
Organizational Affiliation: