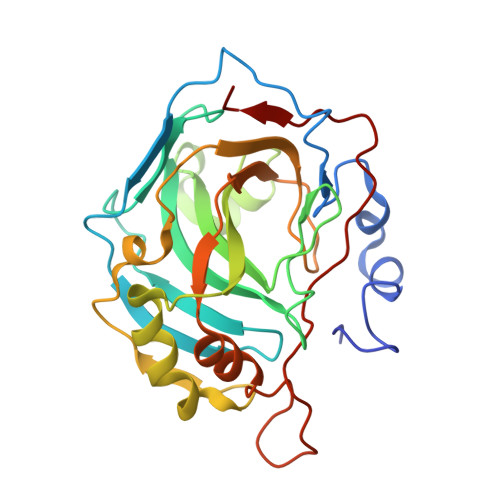
Structure of a 129Xe-cryptophane biosensor complexed with human carbonic anhydrase II.
Aaron, J.A., Chambers, J.M., Jude, K.M., Di Costanzo, L., Dmochowski, I.J., Christianson, D.W.(2008) J Am Chem Soc 130: 6942-6943
- PubMed: 18461940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja802214x
- Primary Citation Related Structures:
3CYU - PubMed Abstract:
Cryptophanes represent an exciting class of xenon-encapsulating molecules that can be exploited as probes for nuclear magnetic resonance imaging. The 1.70 A resolution crystal structure of a cryptophane-derivatized benezenesulfonamide complexed with human carbonic anhydrase II shows how an encapsulated xenon atom can be directed to a specific biological target. The crystal structure confirms binding measurements indicating that the cryptophane cage does not strongly interact with the surface of the protein, which may enhance the sensitivity of 129Xe NMR spectroscopic measurements in solution.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, Pennsylvania 19104-6323, USA.
Organizational Affiliation: