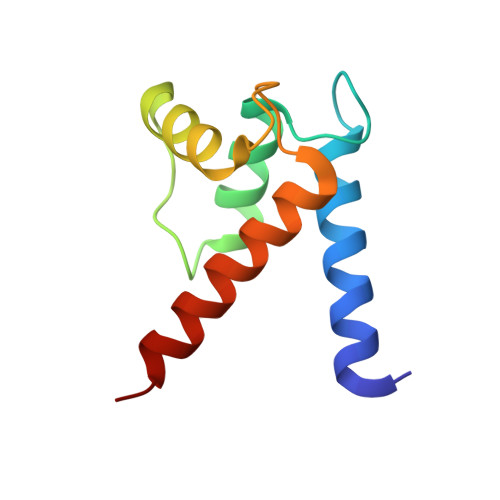
Divalent metal ion complexes of S100B in the absence and presence of pentamidine.
Charpentier, T.H., Wilder, P.T., Liriano, M.A., Varney, K.M., Pozharski, E., MacKerell, A.D., Coop, A., Toth, E.A., Weber, D.J.(2008) J Mol Biology 382: 56-73
- PubMed: 18602402 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.06.047
- Primary Citation Related Structures:
3CR2, 3CR4, 3CR5 - PubMed Abstract:
As part of an effort to inhibit S100B, structures of pentamidine (Pnt) bound to Ca(2+)-loaded and Zn(2+),Ca(2+)-loaded S100B were determined by X-ray crystallography at 2.15 A (R(free)=0.266) and 1.85 A (R(free)=0.243) resolution, respectively. These data were compared to X-ray structures solved in the absence of Pnt, including Ca(2+)-loaded S100B and Zn(2+),Ca(2+)-loaded S100B determined here (1.88 A; R(free)=0.267). In the presence and absence of Zn(2+), electron density corresponding to two Pnt molecules per S100B subunit was mapped for both drug-bound structures. One Pnt binding site (site 1) was adjacent to a p53 peptide binding site on S100B (+/-Zn(2+)), and the second Pnt molecule was mapped to the dimer interface (site 2; +/-Zn(2+)) and in a pocket near residues that define the Zn(2+) binding site on S100B. In addition, a conformational change in S100B was observed upon the addition of Zn(2+) to Ca(2+)-S100B, which changed the conformation and orientation of Pnt bound to sites 1 and 2 of Pnt-Zn(2+),Ca(2+)-S100B when compared to Pnt-Ca(2+)-S100B. That Pnt can adapt to this Zn(2+)-dependent conformational change was unexpected and provides a new mode for S100B inhibition by this drug. These data will be useful for developing novel inhibitors of both Ca(2+)- and Ca(2+),Zn(2+)-bound S100B.
- Department of Biochemistry and Molecular Biology, The University of Maryland School of Medicine, 108 North Greene Street, Baltimore, MD 21201, USA.
Organizational Affiliation: