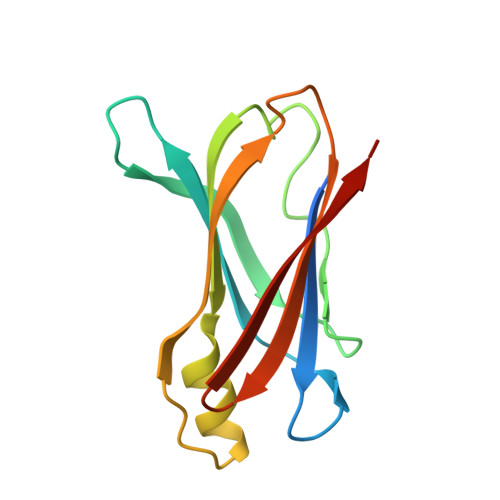
Structural insight into pH-induced conformational changes within the native human transthyretin tetramer.
Palaninathan, S.K., Mohamedmohaideen, N.N., Snee, W.C., Kelly, J.W., Sacchettini, J.C.(2008) J Mol Biology 382: 1157-1167
- PubMed: 18662699 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.07.029
- Primary Citation Related Structures:
3CBR, 3D7P - PubMed Abstract:
Acidification of the transthyretin (TTR) tetramer facilitates dissociation and conformational changes in the protein, allowing alternatively folded monomers to self-assemble into insoluble amyloid fibers by a downhill polymerization mechanism in vitro. To investigate the influence of acidification on the quaternary and tertiary structures of TTR, crystal structures of wild-type human TTR at pH 4.0 and pH 3.5 have been determined to 1.7 A resolution. The acidic pH crystals are isomorphous to most of the previously reported TTR structures, containing two subunits in the asymmetric unit (the so-called A and B subunits) but forming a tetramer through crystallographic symmetry. The pH 4.0 crystal structure reveals that the native fold of the tetramer remains mostly undisturbed. In particular, subunit A of the TTR pH 4.0 structure is very similar to the wild-type TTR pH 7.4 structure with an r.m.s.d. of 0.38 A. In contrast, subunit B of the TTR pH 4.0 structure exhibits several significant changes. The EF-helix (residues 75-81) and the adjacent EF-loop (residues 82-90) show an r.m.s.d. greater than 2.0 A. The acidic residues within this region (Glu72, Asp74, Glu89, and Glu92) undergo significant conformational changes that instigate movement of the EF helix-loop region and make residues Lys70, Lys76, His88, and His90 orient their side chains toward these acidic residues. In particular, Glu89 undergoes a maximum deviation of 5.6 A, occupying Phe87's initial position in the wild-type TTR pH 7.4 structure, and points its side chain into a hydrophobic pocket of the neighboring subunit. In the pH 3.5 structure, the EF helix-loop region is completely disordered. These results demonstrate that acidic conditions increase the susceptibility of the EF helix-loop region of the TTR B subunit to undergo conformational changes and unfold, likely destabilizing the tetramer and identifying at least the initial conformational changes likely occurring within the tetramer that leads to the amyloidogenic monomer.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX 77843, USA.
Organizational Affiliation: