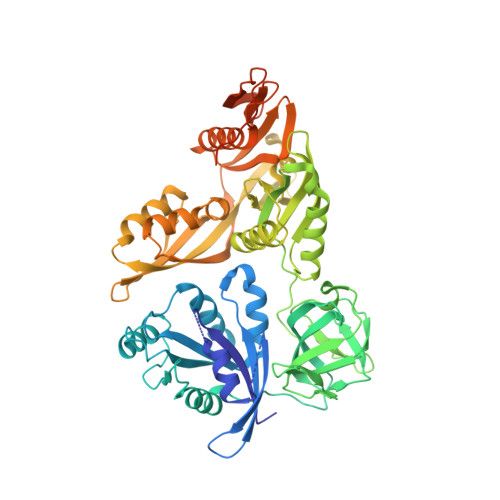
The structure of LepA, the ribosomal back translocase.
Evans, R.N., Blaha, G., Bailey, S., Steitz, T.A.(2008) Proc Natl Acad Sci U S A 105: 4673-4678
- PubMed: 18362332 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0801308105
- Primary Citation Related Structures:
3CB4 - PubMed Abstract:
LepA is a highly conserved elongation factor that promotes the back translocation of tRNAs on the ribosome during the elongation cycle. We have determined the crystal structure of LepA from Escherichia coli at 2.8-A resolution. The high degree of sequence identity between LepA and EF-G is reflected in the structural similarity between the individual homologous domains of LepA and EF-G. However, the orientation of domains III and V in LepA differs from their orientations in EF-G. LepA also contains a C-terminal domain (CTD) not found in EF-G that has a previously unobserved protein fold. The high structural similarity between LepA and EF-G enabled us to derive a homology model for LepA bound to the ribosome using a 7.3-A cryo-EM structure of a complex between EF-G and the 70S ribosome. In this model, the very electrostatically positive CTD of LepA is placed in the direct vicinity of the A site of the large ribosomal subunit, suggesting a possible interaction between the CTD and the back translocated tRNA or 23S rRNA.
- Departments of Molecular Biophysics and Biochemistry and Chemistry and Howard Hughes Medical Institute, Yale University, New Haven, CT 06520-8114, USA.
Organizational Affiliation: