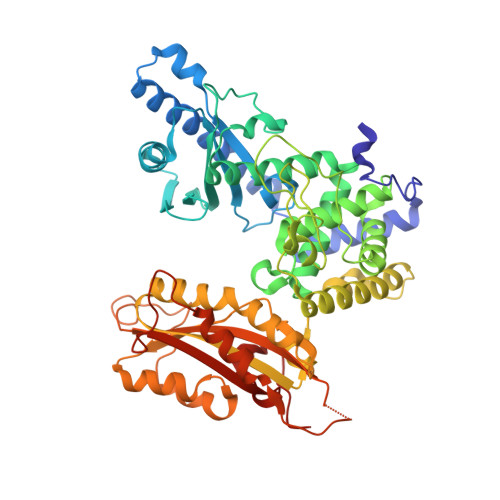
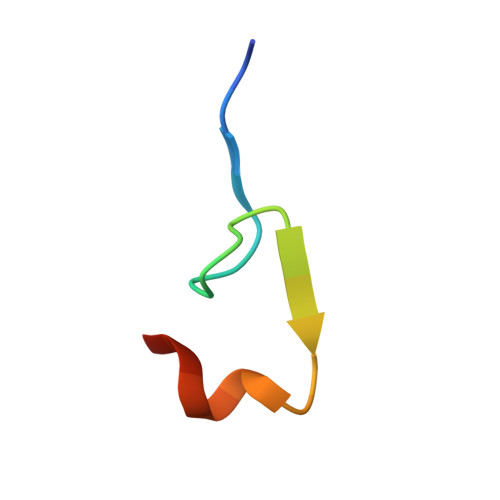
Structure of yeast poly(A) polymerase in complex with a peptide from Fip1, an intrinsically disordered protein.
Meinke, G., Ezeokonkwo, C., Balbo, P., Stafford, W., Moore, C., Bohm, A.(2008) Biochemistry 47: 6859-6869
- PubMed: 18537269 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi800204k
- Primary Citation Related Structures:
3C66 - PubMed Abstract:
In yeast, the mRNA processing enzyme poly(A) polymerase is tethered to the much larger 3'-end processing complex via Fip1, a 36 kDa protein of unknown structure. We report the 2.6 A crystal structure of yeast poly(A) polymerase in complex with a peptide containing residues 80-105 of Fip1. The Fip1 peptide binds to the outside surface of the C-terminal domain of the polymerase. On the basis of this structure, we designed a mutant of the polymerase (V498Y, C485R) that is lethal to yeast. The mutant is unable to bind Fip1 but retains full polymerase activity. Fip1 is found in all eukaryotes and serves to connect poly(A) polymerase to pre-mRNA processing complexes in yeast, plants, and mammals. However, the Fip1 sequence is highly divergent, and residues on both Pap1 and Fip1 at the observed interaction surface are poorly conserved. Herein we demonstrate using analytical ultracentrifugation, circular dichroism, proteolytic studies, and other techniques that, in the absence of Pap1, Fip1 is largely, if not completely, unfolded. We speculate that flexibility may be important for Fip1's function as a molecular scaffold.
- Department of Biochemistry, Tufts University, 136 Harrison Avenue, Boston, Massachusetts 02111, USA.
Organizational Affiliation: