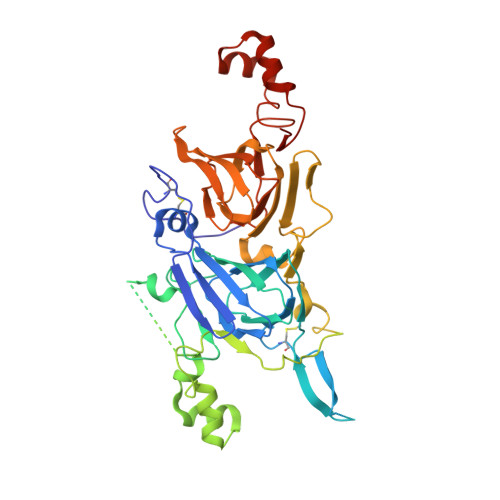
Crystal structure of Ara h 3, a major allergen in peanut.
Jin, T., Guo, F., Chen, Y.W., Howard, A., Zhang, Y.Z.(2009) Mol Immunol 46: 1796-1804
- PubMed: 19251323
- DOI: https://doi.org/10.1016/j.molimm.2009.01.023
- Primary Citation of Related Structures:
3C3V - PubMed Abstract:
The prevalence of food allergy has increased dramatically in recent years. Tremendous research progress has been made in understanding the pathophysiological mechanisms of allergy and in identifying and characterizing food allergens. Peanut is a major food allergen source and Ara h 3 is a major peanut allergen. Using overlapping short peptides, several linear IgE-binding epitopes in Ara h 3 have been defined before. However, the structure of Ara h 3 of the native allergen is not clear and information on conformational epitopes is lacking. Structural characterization of allergens is required for understanding the allergenicity of food allergens and for the development of immunotherapeutic agents. Previously, we have reported the crystallization of Ara h 3 purified from raw peanut. Here we report the crystal structure of Ara h 3 at 1.73A resolution. Mapping of the previously defined linear epitopes on the crystal structure of Ara h 3 indicated that linear epitopes with more solvent exposure were those indicated by the literature to react with more patient sera. The structure of Ara h 3 may be used to assess the importance of conformational epitopes in further investigations.
- Department of Biological, Chemical, and Physical Sciences, Illinois Institute of Technology, Chicago, IL 60616, United States.
Organizational Affiliation: