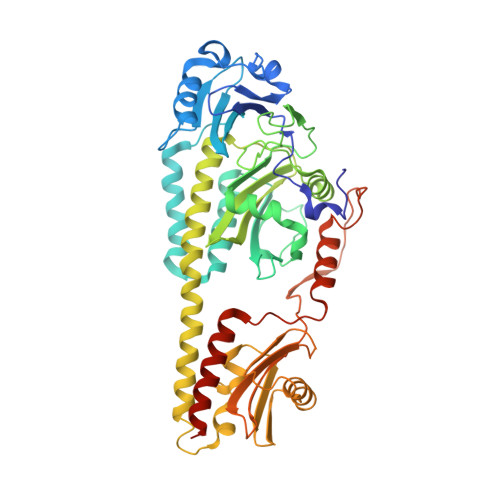
Crystal structure of Pseudomonas aeruginosa bacteriophytochrome: photoconversion and signal transduction.
Yang, X., Kuk, J., Moffat, K.(2008) Proc Natl Acad Sci U S A 105: 14715-14720
- PubMed: 18799746 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0806718105
- Primary Citation Related Structures:
3C2W - PubMed Abstract:
Phytochromes are red-light photoreceptors that regulate light responses in plants, fungi, and bacteria via reversible photoconversion between red (Pr) and far-red (Pfr) light-absorbing states. Here we report the crystal structure at 2.9 A resolution of a bacteriophytochrome from Pseudomonas aeruginosa with an intact, fully photoactive photosensory core domain in its dark-adapted Pfr state. This structure reveals how unusual interdomain interactions, including a knot and an "arm" structure near the chromophore site, bring together the PAS (Per-ARNT-Sim), GAF (cGMP phosphodiesterase/adenyl cyclase/FhlA), and PHY (phytochrome) domains to achieve Pr/Pfr photoconversion. The PAS, GAF, and PHY domains have topologic elements in common and may have a single evolutionary origin. We identify key interactions that stabilize the chromophore in the Pfr state and provide structural and mutational evidence to support the essential role of the PHY domain in efficient Pr/Pfr photoconversion. We also identify a pair of conserved residues that may undergo concerted conformational changes during photoconversion. Modeling of the full-length bacteriophytochrome structure, including its output histidine kinase domain, suggests how local structural changes originating in the photosensory domain modulate interactions between long, cross-domain signaling helices at the dimer interface and are transmitted to the spatially distant effector domain, thereby regulating its histidine kinase activity.
- Department of Biochemistry and Molecular Biology, University of Chicago, Chicago, IL 60637, USA. xiaojingyang@uchicago.edu
Organizational Affiliation: