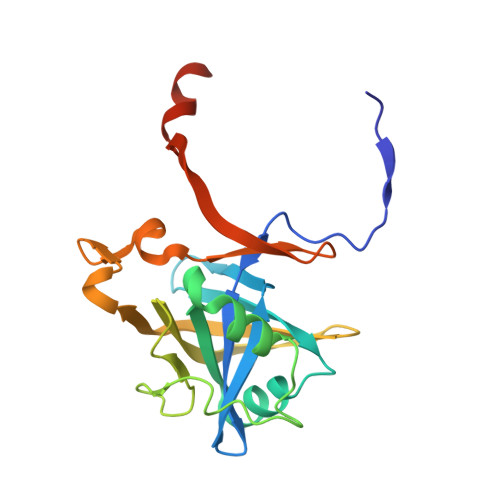
Structural and biochemical characterization of flavoredoxin from the archaeon Methanosarcina acetivorans
Suharti, S., Murakami, K.S., de Vries, S., Ferry, J.G.(2008) Biochemistry 47: 11528-11535
- PubMed: 18842001 Search on PubMed
- DOI: https://doi.org/10.1021/bi801012p
- Primary Citation Related Structures:
3BNK - PubMed Abstract:
Flavoredoxin is a FMN-containing electron transfer protein that functions in the energy-yielding metabolism of Desulfovibrio gigas of the Bacteria domain. Although characterization of this flavoredoxin is the only one reported, a database search revealed homologues widely distributed in both the Bacteria and Archaea domains that define a novel family. To improve our understanding of this family, a flavoredoxin from Methanosarcina acetivorans of the Archaea domain was produced in Escherichia coli and biochemically characterized, and a high-resolution crystal structure was determined. The protein was shown to be a homodimer with a subunit molecular mass of 21 kDa containing one noncovalently bound FMN per monomer. Redox titration showed an E(m) of -271 mV with two electrons, consistent with no semiquinone observed in the potential range studied, a result suggesting the flavoredoxin functions as a two-electron carrier. However, neither of the obligate two-electron carriers, NAD(P)H and coenzyme F420H2, was a competent electron donor, whereas 2[4Fe-4S] ferredoxin reduced the flavoredoxin. The X-ray crystal structure determined at 2.05 A resolution revealed a homodimer containing one FMN per monomer, consistent with the biochemical characterization. The isoalloxazine ring of FMN was shown buried within a narrow groove approximately 10 A from the positively charged protein surface that possibly facilitates interaction with the negatively charged ferredoxin. The structure provides a basis for predicting the mechanism by which electrons are transferred between ferredoxin and FMN. The FMN is bound with hydrogen bonds to the isoalloxazine ring and electrostatic interactions with the phosphate moiety that, together with sequence analyses of homologues, indicate a novel FMN binding motif for the flavoredoxin family.
- Department of Biochemistry and Molecular Biology, The Pennsylvania State University, University Park, Pennsylvania 16802, USA.
Organizational Affiliation: