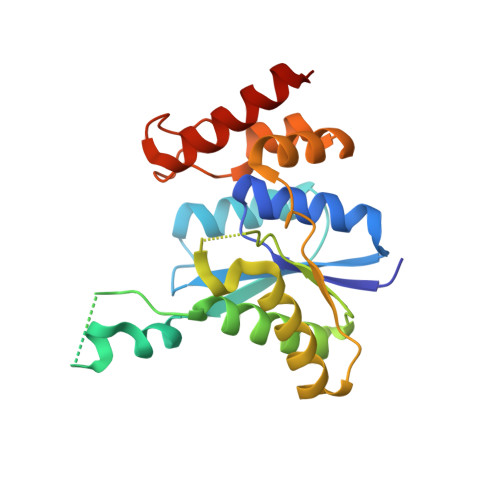
Crystal structure of QueC from Bacillus subtilis: An enzyme involved in preQ(1) biosynthesis.
Cicmil, N., Huang, R.H.(2008) Proteins 72: 1084-1088
- PubMed: 18491386 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22098
- Primary Citation Related Structures:
3BL5 - Department of Biochemistry, University of Illinois at Urbana-Champaign, Urbana, Illinois 61801, USA.
Organizational Affiliation: