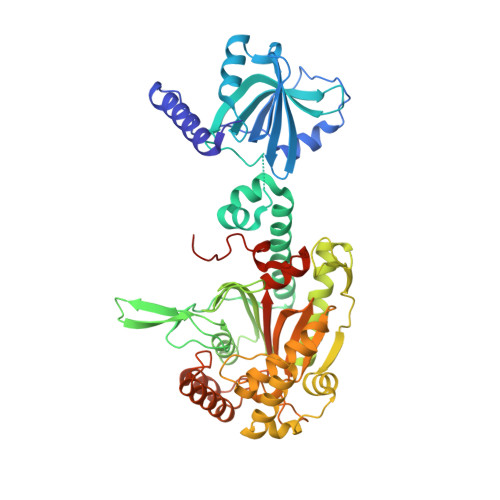
Crystal structure of tetrameric form of human lysyl-tRNA synthetase: Implications for multisynthetase complex formation
Guo, M., Ignatov, M., Musier-Forsyth, K., Schimmel, P., Yang, X.L.(2008) Proc Natl Acad Sci U S A 105: 2331-2336
- PubMed: 18272479 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0712072105
- Primary Citation Related Structures:
3BJU - PubMed Abstract:
In mammals, many aminoacyl-tRNA synthetases are bound together in a multisynthetase complex (MSC) as a reservoir of procytokines and regulation molecules for functions beyond aminoacylation. The alpha(2) homodimeric lysyl-tRNA synthetase (LysRS) is tightly bound in the MSC and, under specific conditions, is secreted to trigger a proinflammatory response. Results by others suggest that alpha(2) LysRS is tightly bound into the core of the MSC with homodimeric beta(2) p38, a scaffolding protein that itself is multifunctional. Not understood is how the two dimeric proteins combine to make a presumptive alpha(2)beta(2) heterotetramer and, in particular, the location of the surfaces on LysRS that would accommodate the p38 interactions. Here we present a 2.3-A crystal structure of a tetrameric form of human LysRS. The relatively loose (as seen in solution) tetramer interface is assembled from two eukaryote-specific sequences, one in the catalytic- and another in the anticodon-binding domain. This same interface is predicted to provide unique determinants for interaction with p38. The analyses suggest how the core of the MSC is assembled and, more generally, that interactions and functions of synthetases can be built and regulated through dynamic protein-protein interfaces. These interfaces are created from small adaptations to what is otherwise a highly conserved (through evolution) polypeptide sequence.
- The Skaggs Institute for Chemical Biology and Department of Molecular Biology, The Scripps Research Institute, BCC-379, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: